The CALIS Procedure
-
Overview

-
Getting Started

-
Syntax
 Classes of Statements in PROC CALISSingle-Group Analysis SyntaxMultiple-Group Multiple-Model Analysis SyntaxPROC CALIS StatementBOUNDS StatementBY StatementCOSAN StatementCOV StatementDETERM StatementEFFPART StatementFACTOR StatementFITINDEX StatementFREQ StatementGROUP StatementLINCON StatementLINEQS StatementLISMOD StatementLMTESTS StatementMATRIX StatementMEAN StatementMODEL StatementMSTRUCT StatementNLINCON StatementNLOPTIONS StatementOUTFILES StatementPARAMETERS StatementPARTIAL StatementPATH StatementPATHDIAGRAM StatementPCOV StatementPVAR StatementRAM StatementREFMODEL StatementRENAMEPARM StatementSAS Programming StatementsSIMTESTS StatementSTD StatementSTRUCTEQ StatementTESTFUNC StatementVAR StatementVARIANCE StatementVARNAMES StatementWEIGHT Statement
Classes of Statements in PROC CALISSingle-Group Analysis SyntaxMultiple-Group Multiple-Model Analysis SyntaxPROC CALIS StatementBOUNDS StatementBY StatementCOSAN StatementCOV StatementDETERM StatementEFFPART StatementFACTOR StatementFITINDEX StatementFREQ StatementGROUP StatementLINCON StatementLINEQS StatementLISMOD StatementLMTESTS StatementMATRIX StatementMEAN StatementMODEL StatementMSTRUCT StatementNLINCON StatementNLOPTIONS StatementOUTFILES StatementPARAMETERS StatementPARTIAL StatementPATH StatementPATHDIAGRAM StatementPCOV StatementPVAR StatementRAM StatementREFMODEL StatementRENAMEPARM StatementSAS Programming StatementsSIMTESTS StatementSTD StatementSTRUCTEQ StatementTESTFUNC StatementVAR StatementVARIANCE StatementVARNAMES StatementWEIGHT Statement -
Details
 Input Data SetsOutput Data SetsDefault Analysis Type and Default ParameterizationThe COSAN ModelThe FACTOR ModelThe LINEQS ModelThe LISMOD Model and SubmodelsThe MSTRUCT ModelThe PATH ModelThe RAM ModelNaming Variables and ParametersSetting Constraints on ParametersAutomatic Variable SelectionPath Diagrams: Layout Algorithms, Default Settings, and CustomizationEstimation CriteriaRelationships among Estimation CriteriaGradient, Hessian, Information Matrix, and Approximate Standard ErrorsCounting the Degrees of FreedomAssessment of FitCase-Level Residuals, Outliers, Leverage Observations, and Residual DiagnosticsLatent Variable ScoresTotal, Direct, and Indirect EffectsStandardized SolutionsModification IndicesMissing Values and the Analysis of Missing PatternsMeasures of Multivariate KurtosisInitial EstimatesUse of Optimization TechniquesComputational ProblemsDisplayed OutputODS Table NamesODS Graphics
Input Data SetsOutput Data SetsDefault Analysis Type and Default ParameterizationThe COSAN ModelThe FACTOR ModelThe LINEQS ModelThe LISMOD Model and SubmodelsThe MSTRUCT ModelThe PATH ModelThe RAM ModelNaming Variables and ParametersSetting Constraints on ParametersAutomatic Variable SelectionPath Diagrams: Layout Algorithms, Default Settings, and CustomizationEstimation CriteriaRelationships among Estimation CriteriaGradient, Hessian, Information Matrix, and Approximate Standard ErrorsCounting the Degrees of FreedomAssessment of FitCase-Level Residuals, Outliers, Leverage Observations, and Residual DiagnosticsLatent Variable ScoresTotal, Direct, and Indirect EffectsStandardized SolutionsModification IndicesMissing Values and the Analysis of Missing PatternsMeasures of Multivariate KurtosisInitial EstimatesUse of Optimization TechniquesComputational ProblemsDisplayed OutputODS Table NamesODS Graphics -
Examples
 Estimating Covariances and CorrelationsEstimating Covariances and Means SimultaneouslyTesting Uncorrelatedness of VariablesTesting Covariance PatternsTesting Some Standard Covariance Pattern HypothesesLinear Regression ModelMultivariate Regression ModelsMeasurement Error ModelsTesting Specific Measurement Error ModelsMeasurement Error Models with Multiple PredictorsMeasurement Error Models Specified As Linear EquationsConfirmatory Factor ModelsConfirmatory Factor Models: Some VariationsResidual Diagnostics and Robust EstimationThe Full Information Maximum Likelihood MethodComparing the ML and FIML EstimationPath Analysis: Stability of AlienationSimultaneous Equations with Mean Structures and Reciprocal PathsFitting Direct Covariance StructuresConfirmatory Factor Analysis: Cognitive AbilitiesTesting Equality of Two Covariance Matrices Using a Multiple-Group AnalysisTesting Equality of Covariance and Mean Matrices between Independent GroupsIllustrating Various General Modeling LanguagesTesting Competing Path Models for the Career Aspiration DataFitting a Latent Growth Curve ModelHigher-Order and Hierarchical Factor ModelsLinear Relations among Factor LoadingsMultiple-Group Model for Purchasing BehaviorFitting the RAM and EQS Models by the COSAN Modeling LanguageSecond-Order Confirmatory Factor AnalysisLinear Relations among Factor Loadings: COSAN Model SpecificationOrdinal Relations among Factor LoadingsLongitudinal Factor Analysis
Estimating Covariances and CorrelationsEstimating Covariances and Means SimultaneouslyTesting Uncorrelatedness of VariablesTesting Covariance PatternsTesting Some Standard Covariance Pattern HypothesesLinear Regression ModelMultivariate Regression ModelsMeasurement Error ModelsTesting Specific Measurement Error ModelsMeasurement Error Models with Multiple PredictorsMeasurement Error Models Specified As Linear EquationsConfirmatory Factor ModelsConfirmatory Factor Models: Some VariationsResidual Diagnostics and Robust EstimationThe Full Information Maximum Likelihood MethodComparing the ML and FIML EstimationPath Analysis: Stability of AlienationSimultaneous Equations with Mean Structures and Reciprocal PathsFitting Direct Covariance StructuresConfirmatory Factor Analysis: Cognitive AbilitiesTesting Equality of Two Covariance Matrices Using a Multiple-Group AnalysisTesting Equality of Covariance and Mean Matrices between Independent GroupsIllustrating Various General Modeling LanguagesTesting Competing Path Models for the Career Aspiration DataFitting a Latent Growth Curve ModelHigher-Order and Hierarchical Factor ModelsLinear Relations among Factor LoadingsMultiple-Group Model for Purchasing BehaviorFitting the RAM and EQS Models by the COSAN Modeling LanguageSecond-Order Confirmatory Factor AnalysisLinear Relations among Factor Loadings: COSAN Model SpecificationOrdinal Relations among Factor LoadingsLongitudinal Factor Analysis - References
Example 29.20 Confirmatory Factor Analysis: Cognitive Abilities
In this example, cognitive abilities of 64 students from a middle school were measured. The fictitious data contain nine cognitive test scores. Three of the scores were for reading skills, three others were for math skills, and the remaining three were for writing skills. The covariance matrix for the nine variables was obtained. A confirmatory factor analysis with three factors was conducted. The following is the input data set:
title "Confirmatory Factor Analysis Using the FACTOR Modeling Language";
title2 "Cognitive Data";
data cognitive1(type=cov);
_type_='cov';
input _name_ $ reading1 reading2 reading3 math1 math2 math3
writing1 writing2 writing3;
datalines;
reading1 83.024 . . . . . . . .
reading2 50.924 108.243 . . . . . . .
reading3 62.205 72.050 99.341 . . . . . .
math1 22.522 22.474 25.731 82.214 . . . . .
math2 14.157 22.487 18.334 64.423 96.125 . . . .
math3 22.252 20.645 23.214 49.287 58.177 88.625 . . .
writing1 33.433 42.474 41.731 25.318 14.254 27.370 90.734 . .
writing2 24.147 20.487 18.034 22.106 26.105 22.346 53.891 96.543 .
writing3 13.340 20.645 23.314 19.387 28.177 38.635 55.347 52.999 98.445
;
Confirmatory Factor Model with Uncorrelated Factors
You first fit a confirmatory factor model with uncorrelated factors to the data, as shown in the following statements:
proc calis data=cognitive1 nobs=64 modification;
factor
Read_Factor ===> reading1-reading3 ,
Math_Factor ===> math1-math3 ,
Write_Factor ===> writing1-writing3 ;
pvar
Read_Factor Math_Factor Write_Factor = 3 * 1.;
cov
Read_Factor Math_Factor Write_Factor = 3 * 0.;
run;
In the PROC CALIS statement, the number of observations is specified with the NOBS= option. With the MODIFICATION in the PROC CALIS statement, LM (Lagrange Multiplier) tests are conducted. The results of LM tests can suggest the inclusion of additional parameters for a better model fit.
The FACTOR modeling language is most handy when you specify confirmatory factor models. You use the FACTOR
statement to invoke the FACTOR modeling language. Entries in the FACTOR
statement are for specifying factor-variables relationships and are separated by commas. In each entry, you first specify
a latent factor, followed by the right arrow sign ===> (you can use >, =>, ==>, or ===>). Then you specify the observed variables that have nonzero loadings on the factor. For example, in the first entry of FACTOR
statement, you specify that latent factor Read_Factor has nonzero loadings (free parameters) on variables reading1–reading3. Optionally, you can specify the parameter list after you specify the factor-variable relationships. For example, you can
name the loading parameters as in the following specification:
factor Read_Factor ===> reading1-reading3 = load1-load3;
This way, you name the factor loadings with parameter names load1, load2, and load3, respectively. However, in the current example, because the loading parameters are all unconstrained, you can just let PROC
CALIS to generate the parameter names for you. In this example, there are three factors: Read_Factor, Math_Factor, and Write_Factor. These factors have simple cluster structures with the nine observed variables. Each observed variable has only one loading
on exactly one factor.
In the PVAR statement, you can specify the variances of the factors and the error variances of the observed variables. The factor variances in this model are all fixed at 1.0 for identification purposes. You do not need to specify the error variances of the observed variables in the current model because PROC CALIS assumes these are free parameters by default.
In the COV statement, you specify that the covariances among the factors are fixed zeros. There are three covariances among
the three latent factors and therefore you put 3 * 0. for their fixed values. This means that the factors in the current model are uncorrelated. Note that you must specify uncorrelated
factors explicitly in the COV statement because all latent factors are correlated by default.
In Output 29.20.1, the initial model specification is echoed in matrix form. The observed variables and factors are also displayed.
Output 29.20.1: Uncorrelated Factor Model Specification
In the table for initial factor loading matrix, the nine loading parameters are shown to have simple cluster relations with the factors. In the table for initial factor covariance matrix, the diagonal matrix shows that the factors are not correlated. The diagonal elements are fixed at ones so that this matrix is also a correlation matrix for the factors. In the table for initial error variances, the nine variance parameters are shown. As described previously, these error variances are generated by PROC CALIS as default parameters.
In Output 29.20.2, initial estimates are generated by the instrumental variable method and the McDonald method.
Output 29.20.2: Optimization of the Uncorrelated Factor Model: Initial Estimates
| Optimization Start Parameter Estimates |
|||
|---|---|---|---|
| N | Parameter | Estimate | Gradient |
| 1 | _Parm1 | 7.15372 | 0.00851 |
| 2 | _Parm2 | 7.80225 | -0.00170 |
| 3 | _Parm3 | 8.70856 | -0.00602 |
| 4 | _Parm4 | 7.68637 | 0.00272 |
| 5 | _Parm5 | 8.01765 | -0.01096 |
| 6 | _Parm6 | 7.05012 | 0.00932 |
| 7 | _Parm7 | 8.76776 | -0.0009955 |
| 8 | _Parm8 | 5.96161 | -0.01335 |
| 9 | _Parm9 | 7.23168 | 0.01665 |
| 10 | _Add1 | 31.84831 | -0.00179 |
| 11 | _Add2 | 47.36790 | 0.0003461 |
| 12 | _Add3 | 23.50199 | 0.00257 |
| 13 | _Add4 | 23.13374 | -0.0008384 |
| 14 | _Add5 | 31.84224 | 0.00280 |
| 15 | _Add6 | 38.92075 | -0.00167 |
| 16 | _Add7 | 13.86035 | -0.00579 |
| 17 | _Add8 | 61.00217 | 0.00115 |
| 18 | _Add9 | 46.14784 | -0.00300 |
| Value of Objective Function = 0.9103815918 | |||
These initial estimates turn out to be pretty good, in the sense that only three more iterations are needed to converge to the maximum likelihood estimates and the final function value 0.784 does not change much from the initial function value 0.910, as shown in Output 29.20.3.
Output 29.20.3: Optimization of the Uncorrelated Factor Model: Iteration Summary
The fit summary is shown in Output 29.20.4.
Output 29.20.4: Fit of the Uncorrelated Factor Model
| Fit Summary | ||
|---|---|---|
| Modeling Info | Number of Observations | 64 |
| Number of Variables | 9 | |
| Number of Moments | 45 | |
| Number of Parameters | 18 | |
| Number of Active Constraints | 0 | |
| Baseline Model Function Value | 4.3182 | |
| Baseline Model Chi-Square | 272.0467 | |
| Baseline Model Chi-Square DF | 36 | |
| Pr > Baseline Model Chi-Square | <.0001 | |
| Absolute Index | Fit Function | 0.7837 |
| Chi-Square | 49.3752 | |
| Chi-Square DF | 27 | |
| Pr > Chi-Square | 0.0054 | |
| Z-Test of Wilson & Hilferty | 2.5474 | |
| Hoelter Critical N | 52 | |
| Root Mean Square Residual (RMR) | 19.5739 | |
| Standardized RMR (SRMR) | 0.2098 | |
| Goodness of Fit Index (GFI) | 0.8555 | |
| Parsimony Index | Adjusted GFI (AGFI) | 0.7592 |
| Parsimonious GFI | 0.6416 | |
| RMSEA Estimate | 0.1147 | |
| RMSEA Lower 90% Confidence Limit | 0.0617 | |
| RMSEA Upper 90% Confidence Limit | 0.1646 | |
| Probability of Close Fit | 0.0271 | |
| ECVI Estimate | 1.4630 | |
| ECVI Lower 90% Confidence Limit | 1.2069 | |
| ECVI Upper 90% Confidence Limit | 1.8687 | |
| Akaike Information Criterion | 85.3752 | |
| Bozdogan CAIC | 142.2351 | |
| Schwarz Bayesian Criterion | 124.2351 | |
| McDonald Centrality | 0.8396 | |
| Incremental Index | Bentler Comparative Fit Index | 0.9052 |
| Bentler-Bonett NFI | 0.8185 | |
| Bentler-Bonett Non-normed Index | 0.8736 | |
| Bollen Normed Index Rho1 | 0.7580 | |
| Bollen Non-normed Index Delta2 | 0.9087 | |
| James et al. Parsimonious NFI | 0.6139 | |
Using the chi-square model test criterion, the uncorrelated factor model should be rejected at 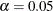 . The RMSEA estimate is 0.1147, which is not indicative of a good fit according to Browne and Cudeck (1993) Other indices might suggest only a marginal good fit. For example, Bentler’s comparative fit index and Bollen nonnormed
index delta2 are both above 0.90. However, many other do not attain this 0.90 level. For example, adjusted GFI is only 0.759.
It is thus safe to conclude that there could be some improvements on the model fit.
. The RMSEA estimate is 0.1147, which is not indicative of a good fit according to Browne and Cudeck (1993) Other indices might suggest only a marginal good fit. For example, Bentler’s comparative fit index and Bollen nonnormed
index delta2 are both above 0.90. However, many other do not attain this 0.90 level. For example, adjusted GFI is only 0.759.
It is thus safe to conclude that there could be some improvements on the model fit.
The MODIFICATION option in the PROC CALIS statement has been used to request for computing the LM test indices for model modifications. The results are shown in Output 29.20.5.
Output 29.20.5: Lagrange Multiplier Tests
| Rank Order of the 10 Largest LM Stat for Factor Loadings | ||||
|---|---|---|---|---|
| Variable | Factor | LM Stat | Pr > ChiSq | Parm Change |
| writing1 | Read_Factor | 9.76596 | 0.0018 | 2.95010 |
| math3 | Write_Factor | 3.58077 | 0.0585 | 1.89703 |
| math1 | Read_Factor | 2.15312 | 0.1423 | 1.17976 |
| writing3 | Math_Factor | 1.87637 | 0.1707 | 1.41298 |
| math3 | Read_Factor | 1.02954 | 0.3103 | 0.95427 |
| reading2 | Write_Factor | 0.91230 | 0.3395 | 0.99933 |
| writing2 | Math_Factor | 0.86221 | 0.3531 | 0.95672 |
| reading1 | Write_Factor | 0.63403 | 0.4259 | 0.73916 |
| math1 | Write_Factor | 0.55602 | 0.4559 | 0.63906 |
| reading2 | Math_Factor | 0.55362 | 0.4568 | 0.74628 |
| Rank Order of the 10 Largest LM Stat for Error Variances and Covariances | ||||
|---|---|---|---|---|
| Error of |
Error of |
LM Stat | Pr > ChiSq | Parm Change |
| writing1 | math2 | 5.45986 | 0.0195 | -13.16822 |
| writing1 | math1 | 5.05573 | 0.0245 | 12.32431 |
| writing3 | math3 | 3.93014 | 0.0474 | 13.59149 |
| writing3 | math1 | 2.83209 | 0.0924 | -9.86342 |
| writing2 | reading1 | 2.56677 | 0.1091 | 10.15901 |
| writing2 | math2 | 1.94879 | 0.1627 | 8.40273 |
| writing2 | reading3 | 1.75181 | 0.1856 | -7.82777 |
| writing3 | reading1 | 1.57978 | 0.2088 | -7.97915 |
| writing1 | reading2 | 1.34894 | 0.2455 | 7.77158 |
| writing2 | math3 | 1.11704 | 0.2906 | -7.23762 |
Three different tables for ranking the LM test results are shown. In the first table, the new loading parameters that would
improve the model fit the most are shown first. For example, in the first row a new factor loading of writing1 on the Read_Factor is suggested to improve the model fit the most. The 'LM Stat' value is 9.77. This is an approximation of the chi-square drop
if this parameter was included in the model. The 'Pr > ChiSq' value of 0.0018 indicates a significant improvement of model
fit at 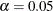 . Nine more new loading parameters are suggested in the table, with less and less statistical significance in the change of
model fit chi-square. Note that these approximate chi-squares are one-at-a-time chi-square changes. That means that the overall
chi-square drop is not a simple sum of individual chi-square changes when you include two or more new parameters in the modified
model.
. Nine more new loading parameters are suggested in the table, with less and less statistical significance in the change of
model fit chi-square. Note that these approximate chi-squares are one-at-a-time chi-square changes. That means that the overall
chi-square drop is not a simple sum of individual chi-square changes when you include two or more new parameters in the modified
model.
The other two tables in Output 29.20.5 shows the new parameters in factor covariances, error variances, or error covariances that would result in a better model fit. The table for the new parameters of the factor covariance matrix indicates that adding each of the covariances among factors might lead to a statistically significant improvement in model fit. The largest 'LM Stat' value in this table is 8.95, which is smaller than that of the largest 'LM Stat' for the factor loading parameters. Despite this, it is more reasonable to add the covariance parameters among factors first to determine whether that improves the model fit.
Confirmatory Factor Model with Correlated Factors
To fit the corresponding confirmatory factor model with correlated factors, you can remove the fixed zeros from the COV statement in the preceding specification, as shown in the following statements:
proc calis data=cognitive1 nobs=64 modification;
factor
Read_Factor ===> reading1-reading3 ,
Math_Factor ===> math1-math3 ,
Write_Factor ===> writing1-writing3 ;
pvar
Read_Factor Math_Factor Write_Factor = 3 * 1.;
cov
Read_Factor Math_Factor Write_Factor /* = 3 * 0. */;
run;
In the COV statement, you comment out the fixed zeros so that the covariances among the latent factors are now free parameters. An alternative way is to delete the entire COV statement so that the covariances among factors are free parameters by the FACTOR model default.
The fit summary of the correlated factor model is shown in Output 29.20.6.
Output 29.20.6: Fit of the Correlated Factor Model
| Fit Summary | ||
|---|---|---|
| Modeling Info | Number of Observations | 64 |
| Number of Variables | 9 | |
| Number of Moments | 45 | |
| Number of Parameters | 21 | |
| Number of Active Constraints | 0 | |
| Baseline Model Function Value | 4.3182 | |
| Baseline Model Chi-Square | 272.0467 | |
| Baseline Model Chi-Square DF | 36 | |
| Pr > Baseline Model Chi-Square | <.0001 | |
| Absolute Index | Fit Function | 0.4677 |
| Chi-Square | 29.4667 | |
| Chi-Square DF | 24 | |
| Pr > Chi-Square | 0.2031 | |
| Z-Test of Wilson & Hilferty | 0.8320 | |
| Hoelter Critical N | 78 | |
| Root Mean Square Residual (RMR) | 5.7038 | |
| Standardized RMR (SRMR) | 0.0607 | |
| Goodness of Fit Index (GFI) | 0.9109 | |
| Parsimony Index | Adjusted GFI (AGFI) | 0.8330 |
| Parsimonious GFI | 0.6073 | |
| RMSEA Estimate | 0.0601 | |
| RMSEA Lower 90% Confidence Limit | 0.0000 | |
| RMSEA Upper 90% Confidence Limit | 0.1244 | |
| Probability of Close Fit | 0.3814 | |
| ECVI Estimate | 1.2602 | |
| ECVI Lower 90% Confidence Limit | 1.2453 | |
| ECVI Upper 90% Confidence Limit | 1.5637 | |
| Akaike Information Criterion | 71.4667 | |
| Bozdogan CAIC | 137.8032 | |
| Schwarz Bayesian Criterion | 116.8032 | |
| McDonald Centrality | 0.9582 | |
| Incremental Index | Bentler Comparative Fit Index | 0.9768 |
| Bentler-Bonett NFI | 0.8917 | |
| Bentler-Bonett Non-normed Index | 0.9653 | |
| Bollen Normed Index Rho1 | 0.8375 | |
| Bollen Non-normed Index Delta2 | 0.9780 | |
| James et al. Parsimonious NFI | 0.5945 | |
The model fit chi-square value is 29.47, which is about 20 less than the model with uncorrelated factors. The p-value is 0.20, indicating a satisfactory model fit. The RMSEA value is 0.06, which is close to 0.05, a value recommended as an indication of good model fit by Browne and Cudeck (1993) More fit indices that do not attain the 0.9 level with the uncorrelated factor model now have values close to or above 0.9. These include the goodness-of-fit index (GFI), McDonald centrality, Bentler-Bonnet NFI, and Bentler-Bonnet nonnormed index. By all counts, the correlated factor model is a much better fit than the uncorrelated factor model.
In Output 29.20.7, the estimation results for factor loadings are shown. All these loadings are statistically significant, indicating non-chance relationships with the factors.
Output 29.20.7: Estimation of the Factor Loading Matrix
| Factor Loading Matrix: Estimate/StdErr/t-value/p-value | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Read_Factor | Math_Factor | Write_Factor | ||||||||||||||||
| reading1 |
|
|
|
|||||||||||||||
| reading2 |
|
|
|
|||||||||||||||
| reading3 |
|
|
|
|||||||||||||||
| math1 |
|
|
|
|||||||||||||||
| math2 |
|
|
|
|||||||||||||||
| math3 |
|
|
|
|||||||||||||||
| writing1 |
|
|
|
|||||||||||||||
| writing2 |
|
|
|
|||||||||||||||
| writing3 |
|
|
|
|||||||||||||||
In Output 29.20.8, the factor covariance matrix is shown. Because the diagonal elements are all ones, the off-diagonal elements are correlations among factors. The correlations range from 0.30–0.5. These factors are moderately correlated.
Output 29.20.8: Estimation of the Correlations of Factors
| Factor Covariance Matrix: Estimate/StdErr/t-value/p-value | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Read_Factor | Math_Factor | Write_Factor | ||||||||||||||||
| Read_Factor |
|
|
|
|||||||||||||||
| Math_Factor |
|
|
|
|||||||||||||||
| Write_Factor |
|
|
|
|||||||||||||||
In Output 29.20.9, the error variances for variables are shown.
Output 29.20.9: Estimation of the Error Variances
| Error Variances | |||||
|---|---|---|---|---|---|
| Variable | Parameter | Estimate | Standard Error |
t Value | Pr > |t| |
| reading1 | _Add1 | 37.24939 | 8.33997 | 4.4664 | <.0001 |
| reading2 | _Add2 | 46.49695 | 10.69869 | 4.3460 | <.0001 |
| reading3 | _Add3 | 15.90447 | 9.26097 | 1.7174 | 0.0859 |
| math1 | _Add4 | 25.22889 | 7.72269 | 3.2669 | 0.0011 |
| math2 | _Add5 | 24.89032 | 8.98327 | 2.7707 | 0.0056 |
| math3 | _Add6 | 42.12110 | 9.20362 | 4.5766 | <.0001 |
| writing1 | _Add7 | 27.24965 | 10.36489 | 2.6290 | 0.0086 |
| writing2 | _Add8 | 49.28881 | 11.39812 | 4.3243 | <.0001 |
| writing3 | _Add9 | 48.10684 | 11.48868 | 4.1873 | <.0001 |
All t values except the one for reading3 are greater than 2, a value close to a critical t value at 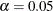 . This means that the error variance for
. This means that the error variance for reading3 could have been zero in the population, or it could have been nonzero but the current sample just has this insignificant
value by chance (that is, a Type 2 error). Further research is needed to confirm either way.
In addition to the parameter estimation results, PROC CALIS also outputs supplementary results that could be useful for interpretations. In Output 29.20.10, the squared multiple correlations and the factor scores regression coefficients are shown.
Output 29.20.10: Supplementary Estimation Results
| Squared Multiple Correlations | |||
|---|---|---|---|
| Variable | Error Variance | Total Variance | R-Square |
| reading1 | 37.24939 | 83.02400 | 0.5513 |
| reading2 | 46.49695 | 108.24300 | 0.5704 |
| reading3 | 15.90447 | 99.34100 | 0.8399 |
| math1 | 25.22889 | 82.21400 | 0.6931 |
| math2 | 24.89032 | 96.12500 | 0.7411 |
| math3 | 42.12110 | 88.62500 | 0.5247 |
| writing1 | 27.24965 | 90.73400 | 0.6997 |
| writing2 | 49.28881 | 96.54300 | 0.4895 |
| writing3 | 48.10684 | 98.44500 | 0.5113 |
| Factor Scores Regression Coefficients | |||
|---|---|---|---|
| Read_Factor | Math_Factor | Write_Factor | |
| reading1 | 0.0200 | 0.000681 | 0.001985 |
| reading2 | 0.0186 | 0.000633 | 0.001847 |
| reading3 | 0.0633 | 0.002152 | 0.006275 |
| math1 | 0.001121 | 0.0403 | 0.002808 |
| math2 | 0.001271 | 0.0457 | 0.003183 |
| math3 | 0.000607 | 0.0218 | 0.001520 |
| writing1 | 0.003195 | 0.002744 | 0.0513 |
| writing2 | 0.001524 | 0.001309 | 0.0245 |
| writing3 | 0.001611 | 0.001384 | 0.0259 |
The percentages of variance for the observed variables that can be explained by the factors are shown in the 'R-Square' column of the table for squared multiple correlations (R-squares). These R-squares can be interpreted meaningfully because there is no reciprocal relationships among variables or correlated errors in the model. All estimates of R-squares are bounded between 0 and 1.
In the table for factor scores regression coefficients, entries are coefficients for the variables you can use to create the
factor scores. The larger the coefficient, the more influence of the corresponding variable for creating the factor scores.
It makes intuitive sense to see the cluster pattern of these coefficients—the reading measures are more important to create
the latent variable scores of Read_Factor and so on.