The GENMOD Procedure
-
Overview

-
Getting Started

-
Syntax
 PROC GENMOD StatementASSESS StatementBAYES StatementBY StatementCLASS StatementCODE StatementCONTRAST StatementDEVIANCE StatementEFFECTPLOT StatementESTIMATE StatementEXACT StatementEXACTOPTIONS StatementFREQ StatementFWDLINK StatementINVLINK StatementLSMEANS StatementLSMESTIMATE StatementMODEL StatementOUTPUT StatementProgramming StatementsREPEATED StatementSLICE StatementSTORE StatementSTRATA StatementVARIANCE StatementWEIGHT StatementZEROMODEL Statement
PROC GENMOD StatementASSESS StatementBAYES StatementBY StatementCLASS StatementCODE StatementCONTRAST StatementDEVIANCE StatementEFFECTPLOT StatementESTIMATE StatementEXACT StatementEXACTOPTIONS StatementFREQ StatementFWDLINK StatementINVLINK StatementLSMEANS StatementLSMESTIMATE StatementMODEL StatementOUTPUT StatementProgramming StatementsREPEATED StatementSLICE StatementSTORE StatementSTRATA StatementVARIANCE StatementWEIGHT StatementZEROMODEL Statement -
Details
 Generalized Linear Models TheorySpecification of EffectsParameterization Used in PROC GENMODType 1 AnalysisType 3 AnalysisConfidence Intervals for ParametersF StatisticsLagrange Multiplier StatisticsPredicted Values of the MeanResidualsMultinomial ModelsZero-Inflated ModelsTweedie Distribution For Generalized Linear ModelsGeneralized Estimating EquationsAssessment of Models Based on Aggregates of ResidualsCase Deletion Diagnostic StatisticsBayesian AnalysisExact Logistic and Exact Poisson RegressionResponse Level OrderingMissing ValuesDisplayed Output for Classical AnalysisDisplayed Output for Bayesian AnalysisDisplayed Output for Exact AnalysisODS Table NamesODS Graphics
Generalized Linear Models TheorySpecification of EffectsParameterization Used in PROC GENMODType 1 AnalysisType 3 AnalysisConfidence Intervals for ParametersF StatisticsLagrange Multiplier StatisticsPredicted Values of the MeanResidualsMultinomial ModelsZero-Inflated ModelsTweedie Distribution For Generalized Linear ModelsGeneralized Estimating EquationsAssessment of Models Based on Aggregates of ResidualsCase Deletion Diagnostic StatisticsBayesian AnalysisExact Logistic and Exact Poisson RegressionResponse Level OrderingMissing ValuesDisplayed Output for Classical AnalysisDisplayed Output for Bayesian AnalysisDisplayed Output for Exact AnalysisODS Table NamesODS Graphics -
Examples
 Logistic RegressionNormal Regression, Log Link Gamma Distribution Applied to Life DataOrdinal Model for Multinomial DataGEE for Binary Data with Logit Link FunctionLog Odds Ratios and the ALR AlgorithmLog-Linear Model for Count DataModel Assessment of Multiple Regression Using Aggregates of ResidualsAssessment of a Marginal Model for Dependent DataBayesian Analysis of a Poisson Regression ModelExact Poisson RegressionTweedie Regression
Logistic RegressionNormal Regression, Log Link Gamma Distribution Applied to Life DataOrdinal Model for Multinomial DataGEE for Binary Data with Logit Link FunctionLog Odds Ratios and the ALR AlgorithmLog-Linear Model for Count DataModel Assessment of Multiple Regression Using Aggregates of ResidualsAssessment of a Marginal Model for Dependent DataBayesian Analysis of a Poisson Regression ModelExact Poisson RegressionTweedie Regression - References
Example 44.2 Normal Regression, Log Link
Consider the following data, where x is an explanatory variable and y is the response variable. It appears that y varies nonlinearly with x and that the variance is approximately constant. A normal distribution with a log link function is chosen to model these
data; that is, 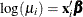 so that
so that 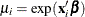 .
.
data nor; input x y; datalines; 0 5 0 7 0 9 1 7 1 10 1 8 2 11 2 9 3 16 3 13 3 14 4 25 4 24 5 34 5 32 5 30 ;
The following SAS statements produce the analysis with the normal distribution and log link:
proc genmod data=nor;
model y = x / dist = normal
link = log;
output out = Residuals
pred = Pred
resraw = Resraw
reschi = Reschi
resdev = Resdev
stdreschi = Stdreschi
stdresdev = Stdresdev
reslik = Reslik;
run;
The OUTPUT statement is specified to produce a data set that contains predicted values and residuals for each observation. This data set can be useful for further analysis, such as residual plotting.
The results from these statements are displayed in Output 44.2.1.
Output 44.2.1: Log-Linked Normal Regression
| Criteria For Assessing Goodness Of Fit | |||
|---|---|---|---|
| Criterion | DF | Value | Value/DF |
| Deviance | 14 | 52.3000 | 3.7357 |
| Scaled Deviance | 14 | 16.0000 | 1.1429 |
| Pearson Chi-Square | 14 | 52.3000 | 3.7357 |
| Scaled Pearson X2 | 14 | 16.0000 | 1.1429 |
| Log Likelihood | -32.1783 | ||
| Full Log Likelihood | -32.1783 | ||
| AIC (smaller is better) | 70.3566 | ||
| AICC (smaller is better) | 72.3566 | ||
| BIC (smaller is better) | 72.6743 | ||
| Analysis Of Maximum Likelihood Parameter Estimates | |||||||
|---|---|---|---|---|---|---|---|
| Parameter | DF | Estimate | Standard Error |
Wald 95% Confidence Limits | Wald Chi-Square | Pr > ChiSq | |
| Intercept | 1 | 1.7214 | 0.0894 | 1.5461 | 1.8966 | 370.76 | <.0001 |
| x | 1 | 0.3496 | 0.0206 | 0.3091 | 0.3901 | 286.64 | <.0001 |
| Scale | 1 | 1.8080 | 0.3196 | 1.2786 | 2.5566 | ||
| Note: | The scale parameter was estimated by maximum likelihood. |
The PROC GENMOD scale parameter, in the case of the normal distribution, is the standard deviation. By default, the scale parameter is estimated by maximum likelihood. You can specify a fixed standard deviation by using the NOSCALE and SCALE= options in the MODEL statement.
proc print data=Residuals; run;
Output 44.2.2: Data Set of Predicted Values and Residuals
| Obs | x | y | Pred | Reschi | Resraw | Resdev | Stdreschi | Stdresdev | Reslik |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 0 | 5 | 5.5921 | -0.59212 | -0.59212 | -0.59212 | -0.34036 | -0.34036 | -0.34036 |
| 2 | 0 | 7 | 5.5921 | 1.40788 | 1.40788 | 1.40788 | 0.80928 | 0.80928 | 0.80928 |
| 3 | 0 | 9 | 5.5921 | 3.40788 | 3.40788 | 3.40788 | 1.95892 | 1.95892 | 1.95892 |
| 4 | 1 | 7 | 7.9324 | -0.93243 | -0.93243 | -0.93243 | -0.54093 | -0.54093 | -0.54093 |
| 5 | 1 | 10 | 7.9324 | 2.06757 | 2.06757 | 2.06757 | 1.19947 | 1.19947 | 1.19947 |
| 6 | 1 | 8 | 7.9324 | 0.06757 | 0.06757 | 0.06757 | 0.03920 | 0.03920 | 0.03920 |
| 7 | 2 | 11 | 11.2522 | -0.25217 | -0.25217 | -0.25217 | -0.14686 | -0.14686 | -0.14686 |
| 8 | 2 | 9 | 11.2522 | -2.25217 | -2.25217 | -2.25217 | -1.31166 | -1.31166 | -1.31166 |
| 9 | 3 | 16 | 15.9612 | 0.03878 | 0.03878 | 0.03878 | 0.02249 | 0.02249 | 0.02249 |
| 10 | 3 | 13 | 15.9612 | -2.96122 | -2.96122 | -2.96122 | -1.71738 | -1.71738 | -1.71738 |
| 11 | 3 | 14 | 15.9612 | -1.96122 | -1.96122 | -1.96122 | -1.13743 | -1.13743 | -1.13743 |
| 12 | 4 | 25 | 22.6410 | 2.35897 | 2.35897 | 2.35897 | 1.37252 | 1.37252 | 1.37252 |
| 13 | 4 | 24 | 22.6410 | 1.35897 | 1.35897 | 1.35897 | 0.79069 | 0.79069 | 0.79069 |
| 14 | 5 | 34 | 32.1163 | 1.88366 | 1.88366 | 1.88366 | 1.22914 | 1.22914 | 1.22914 |
| 15 | 5 | 32 | 32.1163 | -0.11634 | -0.11634 | -0.11634 | -0.07592 | -0.07592 | -0.07592 |
| 16 | 5 | 30 | 32.1163 | -2.11634 | -2.11634 | -2.11634 | -1.38098 | -1.38098 | -1.38098 |
The data set of predicted values and residuals (Output 44.2.2) is created by the OUTPUT statement. You can use the PLOTS= option in the PROC GENMOD statement to create plots of predicted values and residuals. Note that raw, Pearson, and deviance residuals are equal in this example. This is a characteristic of the normal distribution and is not true in general for other distributions.