The LOGISTIC Procedure
- Overview
- Getting Started
-
Syntax
 PROC LOGISTIC Statement BY Statement CLASS Statement CONTRAST Statement EFFECT Statement EFFECTPLOT Statement ESTIMATE Statement EXACT Statement EXACTOPTIONS Statement FREQ Statement LSMEANS Statement LSMESTIMATE Statement MODEL Statement ODDSRATIO Statement OUTPUT Statement ROC Statement ROCCONTRAST Statement SCORE Statement SLICE Statement STORE Statement STRATA Statement TEST Statement UNITS Statement WEIGHT Statement
PROC LOGISTIC Statement BY Statement CLASS Statement CONTRAST Statement EFFECT Statement EFFECTPLOT Statement ESTIMATE Statement EXACT Statement EXACTOPTIONS Statement FREQ Statement LSMEANS Statement LSMESTIMATE Statement MODEL Statement ODDSRATIO Statement OUTPUT Statement ROC Statement ROCCONTRAST Statement SCORE Statement SLICE Statement STORE Statement STRATA Statement TEST Statement UNITS Statement WEIGHT Statement -
Details
 Missing Values Response Level Ordering Link Functions and the Corresponding Distributions Determining Observations for Likelihood Contributions Iterative Algorithms for Model Fitting Convergence Criteria Existence of Maximum Likelihood Estimates Effect-Selection Methods Model Fitting Information Generalized Coefficient of Determination Score Statistics and Tests Confidence Intervals for Parameters Odds Ratio Estimation Rank Correlation of Observed Responses and Predicted Probabilities Linear Predictor, Predicted Probability, and Confidence Limits Classification Table Overdispersion The Hosmer-Lemeshow Goodness-of-Fit Test Receiver Operating Characteristic Curves Testing Linear Hypotheses about the Regression Coefficients Regression Diagnostics Scoring Data Sets Conditional Logistic Regression Exact Conditional Logistic Regression Input and Output Data Sets Computational Resources Displayed Output ODS Table Names ODS Graphics
Missing Values Response Level Ordering Link Functions and the Corresponding Distributions Determining Observations for Likelihood Contributions Iterative Algorithms for Model Fitting Convergence Criteria Existence of Maximum Likelihood Estimates Effect-Selection Methods Model Fitting Information Generalized Coefficient of Determination Score Statistics and Tests Confidence Intervals for Parameters Odds Ratio Estimation Rank Correlation of Observed Responses and Predicted Probabilities Linear Predictor, Predicted Probability, and Confidence Limits Classification Table Overdispersion The Hosmer-Lemeshow Goodness-of-Fit Test Receiver Operating Characteristic Curves Testing Linear Hypotheses about the Regression Coefficients Regression Diagnostics Scoring Data Sets Conditional Logistic Regression Exact Conditional Logistic Regression Input and Output Data Sets Computational Resources Displayed Output ODS Table Names ODS Graphics -
Examples
 Stepwise Logistic Regression and Predicted Values Logistic Modeling with Categorical Predictors Ordinal Logistic Regression Nominal Response Data: Generalized Logits Model Stratified Sampling Logistic Regression Diagnostics ROC Curve, Customized Odds Ratios, Goodness-of-Fit Statistics, R-Square, and Confidence Limits Comparing Receiver Operating Characteristic Curves Goodness-of-Fit Tests and Subpopulations Overdispersion Conditional Logistic Regression for Matched Pairs Data Firth’s Penalized Likelihood Compared with Other Approaches Complementary Log-Log Model for Infection Rates Complementary Log-Log Model for Interval-Censored Survival Times Scoring Data Sets Using the LSMEANS Statement
Stepwise Logistic Regression and Predicted Values Logistic Modeling with Categorical Predictors Ordinal Logistic Regression Nominal Response Data: Generalized Logits Model Stratified Sampling Logistic Regression Diagnostics ROC Curve, Customized Odds Ratios, Goodness-of-Fit Statistics, R-Square, and Confidence Limits Comparing Receiver Operating Characteristic Curves Goodness-of-Fit Tests and Subpopulations Overdispersion Conditional Logistic Regression for Matched Pairs Data Firth’s Penalized Likelihood Compared with Other Approaches Complementary Log-Log Model for Infection Rates Complementary Log-Log Model for Interval-Censored Survival Times Scoring Data Sets Using the LSMEANS Statement - References
Example 53.13 Complementary Log-Log Model for Infection Rates
Antibodies produced in response to an infectious disease like malaria remain in the body after the individual has recovered from the disease. A serological test detects the presence or absence of such antibodies. An individual with such antibodies is called seropositive. In geographic areas where the disease is endemic, the inhabitants are at fairly constant risk of infection. The probability of an individual never having been infected in 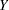 years is
years is 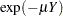 , where
, where 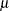 is the mean number of infections per year (see the appendix of Draper, Voller, and Carpenter 1972). Rather than estimating the unknown
is the mean number of infections per year (see the appendix of Draper, Voller, and Carpenter 1972). Rather than estimating the unknown 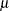 , epidemiologists want to estimate the probability of a person living in the area being infected in one year. This infection rate
, epidemiologists want to estimate the probability of a person living in the area being infected in one year. This infection rate 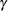 is given by
is given by
 |
The following statements create the data set sero, which contains the results of a serological survey of malarial infection. Individuals of nine age groups (Group) were tested. The variable A represents the midpoint of the age range for each age group. The variable N represents the number of individuals tested in each age group, and the variable R represents the number of individuals that are seropositive.
data sero; input Group A N R; X=log(A); label X='Log of Midpoint of Age Range'; datalines; 1 1.5 123 8 2 4.0 132 6 3 7.5 182 18 4 12.5 140 14 5 17.5 138 20 6 25.0 161 39 7 35.0 133 19 8 47.0 92 25 9 60.0 74 44 ;
For the 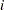 th group with the age midpoint
th group with the age midpoint 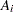 , the probability of being seropositive is
, the probability of being seropositive is  . It follows that
. It follows that
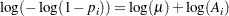 |
By fitting a binomial model with a complementary log-log link function and by using X=log(A) as an offset term, you can estimate 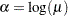 as an intercept parameter. The following statements invoke PROC LOGISTIC to compute the maximum likelihood estimate of
as an intercept parameter. The following statements invoke PROC LOGISTIC to compute the maximum likelihood estimate of  . The LINK=CLOGLOG option is specified to request the complementary log-log link function. Also specified is the CLPARM=PL option, which requests the profile-likelihood confidence limits for
. The LINK=CLOGLOG option is specified to request the complementary log-log link function. Also specified is the CLPARM=PL option, which requests the profile-likelihood confidence limits for  .
.
proc logistic data=sero;
model R/N= / offset=X
link=cloglog
clparm=pl
scale=none;
title 'Constant Risk of Infection';
run;
Results of fitting this constant risk model are shown in Output 53.13.1.
| Constant Risk of Infection |
| Model Information | ||
|---|---|---|
| Data Set | WORK.SERO | |
| Response Variable (Events) | R | |
| Response Variable (Trials) | N | |
| Offset Variable | X | Log of Midpoint of Age Range |
| Model | binary cloglog | |
| Optimization Technique | Fisher's scoring | |
| Number of Observations Read | 9 |
|---|---|
| Number of Observations Used | 9 |
| Sum of Frequencies Read | 1175 |
| Sum of Frequencies Used | 1175 |
| Response Profile | ||
|---|---|---|
| Ordered Value |
Binary Outcome | Total Frequency |
| 1 | Event | 193 |
| 2 | Nonevent | 982 |
| Intercept-Only Model Convergence Status |
|---|
| Convergence criterion (GCONV=1E-8) satisfied. |
| Deviance and Pearson Goodness-of-Fit Statistics | ||||
|---|---|---|---|---|
| Criterion | Value | DF | Value/DF | Pr > ChiSq |
| Deviance | 41.5032 | 8 | 5.1879 | <.0001 |
| Pearson | 50.6883 | 8 | 6.3360 | <.0001 |
| Analysis of Maximum Likelihood Estimates | |||||
|---|---|---|---|---|---|
| Parameter | DF | Estimate | Standard Error |
Wald Chi-Square |
Pr > ChiSq |
| Intercept | 1 | -4.6605 | 0.0725 | 4133.5626 | <.0001 |
| X | 1 | 1.0000 | 0 | . | . |
| Parameter Estimates and Profile-Likelihood Confidence Intervals |
|||
|---|---|---|---|
| Parameter | Estimate | 95% Confidence Limits | |
| Intercept | -4.6605 | -4.8057 | -4.5219 |
Output 53.13.1 shows that the maximum likelihood estimate of 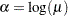 and its estimated standard error are
and its estimated standard error are  and
and  , respectively. The infection rate is estimated as
, respectively. The infection rate is estimated as
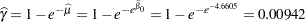 |
The 95% confidence interval for 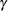 , obtained by back-transforming the 95% confidence interval for
, obtained by back-transforming the 95% confidence interval for  , is (0.0082, 0.0108); that is, there is a 95% chance that, in repeated sampling, the interval of 8 to 11 infections per thousand individuals contains the true infection rate.
, is (0.0082, 0.0108); that is, there is a 95% chance that, in repeated sampling, the interval of 8 to 11 infections per thousand individuals contains the true infection rate.
The goodness-of-fit statistics for the constant risk model are statistically significant (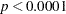 ), indicating that the assumption of constant risk of infection is not correct. You can fit a more extensive model by allowing a separate risk of infection for each age group. Suppose
), indicating that the assumption of constant risk of infection is not correct. You can fit a more extensive model by allowing a separate risk of infection for each age group. Suppose  is the mean number of infections per year for the
is the mean number of infections per year for the 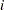 th age group. The probability of seropositive for the
th age group. The probability of seropositive for the 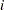 th group with the age midpoint
th group with the age midpoint 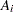 is
is  , so that
, so that
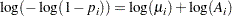 |
In the following statements, a complementary log-log model is fit containing Group as an explanatory classification variable with the GLM coding (so that a dummy variable is created for each age group), no intercept term, and X=log(A) as an offset term. The ODS OUTPUT statement saves the estimates and their 95% profile-likelihood confidence limits to the ClparmPL data set. Note that 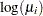 is the regression parameter associated with Group
is the regression parameter associated with Group .
.
proc logistic data=sero;
ods output ClparmPL=ClparmPL;
class Group / param=glm;
model R/N=Group / noint
offset=X
link=cloglog
clparm=pl;
title 'Infectious Rates and 95% Confidence Intervals';
run;
Results of fitting the model with a separate risk of infection are shown in Output 53.13.2.
| Infectious Rates and 95% Confidence Intervals |
| Analysis of Maximum Likelihood Estimates | ||||||
|---|---|---|---|---|---|---|
| Parameter | DF | Estimate | Standard Error |
Wald Chi-Square |
Pr > ChiSq | |
| Group | 1 | 1 | -3.1048 | 0.3536 | 77.0877 | <.0001 |
| Group | 2 | 1 | -4.4542 | 0.4083 | 119.0164 | <.0001 |
| Group | 3 | 1 | -4.2769 | 0.2358 | 328.9593 | <.0001 |
| Group | 4 | 1 | -4.7761 | 0.2674 | 319.0600 | <.0001 |
| Group | 5 | 1 | -4.7165 | 0.2238 | 443.9920 | <.0001 |
| Group | 6 | 1 | -4.5012 | 0.1606 | 785.1350 | <.0001 |
| Group | 7 | 1 | -5.4252 | 0.2296 | 558.1114 | <.0001 |
| Group | 8 | 1 | -4.9987 | 0.2008 | 619.4666 | <.0001 |
| Group | 9 | 1 | -4.1965 | 0.1559 | 724.3157 | <.0001 |
| X | 1 | 1.0000 | 0 | . | . | |
| Parameter Estimates and Profile-Likelihood Confidence Intervals |
||||
|---|---|---|---|---|
| Parameter | Estimate | 95% Confidence Limits | ||
| Group | 1 | -3.1048 | -3.8880 | -2.4833 |
| Group | 2 | -4.4542 | -5.3769 | -3.7478 |
| Group | 3 | -4.2769 | -4.7775 | -3.8477 |
| Group | 4 | -4.7761 | -5.3501 | -4.2940 |
| Group | 5 | -4.7165 | -5.1896 | -4.3075 |
| Group | 6 | -4.5012 | -4.8333 | -4.2019 |
| Group | 7 | -5.4252 | -5.9116 | -5.0063 |
| Group | 8 | -4.9987 | -5.4195 | -4.6289 |
| Group | 9 | -4.1965 | -4.5164 | -3.9037 |
For the first age group (Group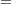 1), the point estimate of
1), the point estimate of 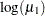 is
is  , which transforms into an infection rate of
, which transforms into an infection rate of  . A 95% confidence interval for this infection rate is obtained by transforming the 95% confidence interval for
. A 95% confidence interval for this infection rate is obtained by transforming the 95% confidence interval for 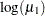 . For the first age group, the lower and upper confidence limits are
. For the first age group, the lower and upper confidence limits are 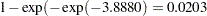 and
and 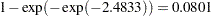 , respectively; that is, there is a 95% chance that, in repeated sampling, the interval of 20 to 80 infections per thousand individuals contains the true infection rate. The following statements perform this transformation on the estimates and confidence limits saved in the ClparmPL data set; the resulting estimated infection rates in one year’s time for each age group are displayed in Table 53.16. Note that the infection rate for the first age group is high compared to that of the other age groups.
, respectively; that is, there is a 95% chance that, in repeated sampling, the interval of 20 to 80 infections per thousand individuals contains the true infection rate. The following statements perform this transformation on the estimates and confidence limits saved in the ClparmPL data set; the resulting estimated infection rates in one year’s time for each age group are displayed in Table 53.16. Note that the infection rate for the first age group is high compared to that of the other age groups.
data ClparmPL; set ClparmPL; Estimate=round( 1000*( 1-exp(-exp(Estimate)) ) ); LowerCL =round( 1000*( 1-exp(-exp(LowerCL )) ) ); UpperCL =round( 1000*( 1-exp(-exp(UpperCL )) ) ); run;
Number Infected per 1,000 People |
|||
|---|---|---|---|
Age |
Point |
95% Confidence Limits |
|
Group |
Estimate |
Lower |
Upper |
1 |
44 |
20 |
80 |
2 |
12 |
5 |
23 |
3 |
14 |
8 |
21 |
4 |
8 |
5 |
14 |
5 |
9 |
6 |
13 |
6 |
11 |
8 |
15 |
7 |
4 |
3 |
7 |
8 |
7 |
4 |
10 |
9 |
15 |
11 |
20 |