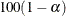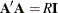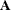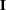-
ALPHA=value
-
sets the confidence level for confidence intervals. The
value of the ALPHA= option must be between 0 and 1, and the default value is 0.05. A confidence level of  produces
produces 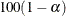 % confidence intervals. The default of ALPHA=0.05 produces 95% confidence intervals.
% confidence intervals. The default of ALPHA=0.05 produces 95% confidence intervals.
-
DATA=SAS-data-set
-
names the SAS data set containing the data to be analyzed.
If you omit the DATA= option, the procedure uses the most recently created SAS data set.
-
INEST=SAS-data-set
-
names the SAS data set that contains initial estimates for
all the parameters in the model. BY-group processing is allowed in setting up the INEST= data set. See the section INEST= Data Set for more information.
-
MISSING
-
treats missing values as a valid (nonmissing)
category for all categorical variables, which include CLASS
, STRATA
, CLUSTER
, and DOMAIN
variables.
By default, if you do not specify the MISSING option, an observation is excluded from the analysis if it has a missing value.
For more information, see the section Missing Values.
-
NAMELEN=n
-
specifies the length of effect names in tables and output
data sets to be n characters, where n is a value between 20 and 200. The default length is 20 characters.
-
NOMCAR
-
requests that the procedure treat missing values
in the variance computation as not missing completely at random (NOMCAR) for Taylor series variance estimation. When you specify the NOMCAR option, PROC SURVEYLOGISTIC computes variance
estimates by analyzing the nonmissing values as a domain or subpopulation, where the entire population includes both nonmissing
and missing domains. See the section Missing Values for more details.
By default, PROC SURVEYLOGISTIC completely excludes an observation from analysis if that observation has a missing value,
unless you specify the MISSING
option. Note that the NOMCAR option has no effect on a classification variable when you specify the MISSING option, which
treats missing values as a valid nonmissing level.
The NOMCAR option applies only to Taylor series variance estimation. The replication methods, which you request with the VARMETHOD=BRR
and VARMETHOD=JACKKNIFE
options, do not use the NOMCAR option.
-
NOSORT
-
suppresses the internal sorting process to shorten
the computation time if the data set is presorted by the STRATA
and CLUSTER
variables. By default, the procedure sorts the data by the STRATA variables if you use the STRATA statement; then the procedure
sorts the data by the CLUSTER variables within strata. If your data are already stored by the order of STRATA and CLUSTER
variables, then you can specify this option to omit this sorting process to reduce the usage of computing resources, especially
when your data set is very large. However, if you specify this NOSORT option while your data are not presorted by STRATA and
CLUSTER variables, then any changes in these variables creates a new stratum or cluster.
-
ORDER=DATA | FORMATTED | FREQ | INTERNAL
-
specifies the sort order for the levels of the response
variable. This option, except for ORDER=FREQ, also determines the sort order for the levels of ClUSTER and DOMAIN variables
and controls STRATA variable levels in the "Stratum Information" table. By default, ORDER=INTERNAL. However, if an ORDER=
option is specified after the response variable, in the MODEL
statement, it overrides this option for the response variable. This option does not affect the ordering of the CLASS variable
levels; see the ORDER=
option in the CLASS
statement for more information.
-
RATE=value | SAS-data-set
R=value | SAS-data-set
-
specifies the sampling rate as a nonnegative
value, or specifies an input data set that contains the stratum sampling rates. The procedure uses this information to compute
a finite population correction for Taylor series variance estimation. The procedure does not use the RATE= option for BRR
or jackknife variance estimation, which you request with the VARMETHOD=BRR
or VARMETHOD=JACKKNIFE
option.
If your sample design has multiple stages, you should specify the first-stage sampling rate,
which is the ratio of the number of PSUs selected to the total number of PSUs in the population.
For a nonstratified sample design, or for a stratified sample design with the same sampling rate in all strata, you should
specify a nonnegative value for the RATE= option. If your design is stratified with different sampling rates in the strata, then you should name a SAS
data set that contains the stratification variables and the sampling rates. See the section Specification of Population Totals and Sampling Rates for more details.
The value in the RATE= option or the values of _RATE_ in the secondary data set must be nonnegative numbers. You can specify value as a number between 0 and 1. Or you can specify value in percentage form as a number between 1 and 100, and PROC SURVEYLOGISTIC converts that number to a proportion. The procedure
treats the value 1 as 100% instead of 1%.
If you do not specify the TOTAL=
or RATE= option, then the Taylor series variance estimation does not include a finite population correction. You cannot specify
both the TOTAL= and RATE= options.
-
TOTAL=value | SAS-data-set
N=value | SAS-data-set
-
specifies the total number of primary sampling units in
the study population as a positive value, or specifies an input data set that contains the stratum population totals. The procedure uses this information to compute
a finite population correction for Taylor series variance estimation. The procedure does not use the TOTAL= option for BRR
or jackknife variance estimation, which you request with the VARMETHOD=BRR
or VARMETHOD=JACKKNIFE
option.
For a nonstratified sample design, or for a stratified sample design with the same population total in all strata, you should
specify a positive value for the TOTAL= option. If your sample design is stratified with different population totals in the strata, then you should
name a SAS data set that contains the stratification variables and the population totals. See the section Specification of Population Totals and Sampling Rates for more details.
If you do not specify the TOTAL= or RATE=
option, then the Taylor series variance estimation does not include a finite population correction. You cannot specify both
the TOTAL= and RATE= options.
-
VARMETHOD=BRR <(method-options)>
VARMETHOD=JACKKNIFE | JK <(method-options)>
VARMETHOD=TAYLOR
-
specifies the variance estimation method. VARMETHOD=TAYLOR requests the Taylor series method, which is the default if you
do not specify the VARMETHOD= option or the REPWEIGHTS statement. VARMETHOD=BRR requests variance estimation by balanced repeated
replication (BRR), and VARMETHOD=JACKKNIFE requests variance estimation by the delete-1 jackknife method.
For VARMETHOD=BRR and VARMETHOD=JACKKNIFE you can specify method-options in parentheses. Table 111.2 summarizes the available method-options.
Table 111.2: Variance Estimation Options
Method-options must be enclosed in parentheses following the method keyword. For example:
varmethod=BRR(reps=60 outweights=myReplicateWeights)
The following values are available for the VARMETHOD= option:
- BRR <(method-options)>
-
requests balanced repeated replication
(BRR) variance estimation. The BRR method requires a stratified sample design with two primary sampling units (PSUs) per
stratum. See the section Balanced Repeated Replication (BRR) Method for more information.
You can specify the following method-options in parentheses following VARMETHOD=BRR:
-
FAY <=value>
-
requests Fay’s method
, a modification of the BRR
method, for variance estimation. See the section Fay’s BRR Method for more information.
You can specify the value of the Fay coefficient, which is used in converting the original sampling weights to replicate weights. The Fay coefficient
must be a nonnegative number less than 1. By default, the value of the Fay coefficient equals 0.5.
-
HADAMARD=SAS-data-set
H=SAS-data-set
-
names a SAS data set that contains the Hadamard matrix
for BRR replicate construction. If you do not provide a Hadamard matrix with the HADAMARD= method-option, PROC SURVEYLOGISTIC generates an appropriate Hadamard matrix for replicate construction. See the sections Balanced Repeated Replication (BRR) Method and Hadamard Matrix for details.
If a Hadamard matrix of a given dimension exists, it is not necessarily unique. Therefore, if you want to use a specific Hadamard
matrix, you must provide the matrix as a SAS data set in the HADAMARD= method-option.
In the HADAMARD= input data set, each variable corresponds to a column of the Hadamard matrix, and each observation corresponds
to a row of the matrix. You can use any variable names in the HADAMARD= data set. All values in the data set must equal either
1 or –1. You must ensure that the matrix you provide is indeed a Hadamard matrix—that is, 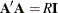 , where
, where 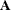 is the Hadamard matrix of dimension R and
is the Hadamard matrix of dimension R and 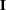 is an identity matrix. PROC SURVEYLOGISTIC does not check the validity of the Hadamard matrix that you provide.
is an identity matrix. PROC SURVEYLOGISTIC does not check the validity of the Hadamard matrix that you provide.
The HADAMARD= input data set must contain at least H variables, where H denotes the number of first-stage strata in your design. If the data set contains more than H variables, the procedure uses only the first H variables. Similarly, the HADAMARD= input data set must contain at least H observations.
If you do not specify the REPS= method-option, then the number of replicates is taken to be the number of observations in the HADAMARD= input data set. If you specify
the number of replicates—for example, REPS=nreps—then the first nreps observations in the HADAMARD= data set are used to construct the replicates.
You can specify the PRINTH option to display the Hadamard matrix that the procedure uses to construct replicates for BRR.
-
OUTWEIGHTS=SAS-data-set
-
names a SAS data set that contains replicate weights. See the section Balanced Repeated Replication (BRR) Method for information about replicate weights. See the section Replicate Weights Output Data Set for more details about the contents of the OUTWEIGHTS= data set.
The OUTWEIGHTS= method-option is not available when you provide replicate weights with the REPWEIGHTS
statement.
-
PRINTH
-
displays the Hadamard matrix.
When you provide your own Hadamard matrix by using the HADAMARD=
method-option, only the rows and columns of the Hadamard matrix that are used by the procedure are displayed. See the sections Balanced Repeated Replication (BRR) Method and Hadamard Matrix for details.
The PRINTH method-option is not available when you provide replicate weights with the REPWEIGHTS
statement because the procedure does not use a Hadamard matrix in this case.
-
REPS=number
-
specifies the number of replicates for BRR variance estimation. The value of number must be an integer greater than 1.
If you do not provide a Hadamard matrix with the HADAMARD=
method-option, the number of replicates should be greater than the number of strata and should be a multiple of 4. See the section Balanced Repeated Replication (BRR) Method for more information. If a Hadamard matrix cannot be constructed for the REPS= value that you specify, the value is increased
until a Hadamard matrix of that dimension can be constructed. Therefore, it is possible for the actual number of replicates
used to be larger than the REPS= value that you specify.
If you provide a Hadamard matrix with the HADAMARD= method-option, the value of REPS= must not be greater than the number of rows in the Hadamard matrix. If you provide a Hadamard matrix
and do not specify the REPS= method-option, the number of replicates equals the number of rows in the Hadamard matrix.
If you do not specify the REPS= or HADAMARD= method-option and do not include a REPWEIGHTS
statement, the number of replicates equals the smallest multiple of 4 that is greater than the number of strata.
If you provide replicate weights with the REPWEIGHTS statement, the procedure does not use the REPS= method-option. With a REPWEIGHTS statement, the number of replicates equals the number of REPWEIGHTS variables.
- JACKKNIFE | JK <(method-options)>
-
requests variance estimation by the delete-1 jackknife method. See the section Jackknife Method for details. If you provide replicate weights with a REPWEIGHTS
statement, VARMETHOD=JACKKNIFE is the default variance estimation method.
You can specify the following method-options in parentheses following VARMETHOD=JACKKNIFE:
- TAYLOR
-
requests Taylor series variance estimation. This is the default method if you do not specify the VARMETHOD= option or a REPWEIGHTS
statement. See the section Taylor Series (Linearization) for more information.
 PROC SURVEYLOGISTIC StatementBY StatementCLASS StatementCLUSTER StatementCONTRAST StatementDOMAIN StatementEFFECT StatementESTIMATE StatementFREQ StatementLSMEANS StatementLSMESTIMATE StatementMODEL StatementOUTPUT StatementREPWEIGHTS StatementSLICE StatementSTORE StatementSTRATA StatementTEST StatementUNITS StatementWEIGHT Statement
PROC SURVEYLOGISTIC StatementBY StatementCLASS StatementCLUSTER StatementCONTRAST StatementDOMAIN StatementEFFECT StatementESTIMATE StatementFREQ StatementLSMEANS StatementLSMESTIMATE StatementMODEL StatementOUTPUT StatementREPWEIGHTS StatementSLICE StatementSTORE StatementSTRATA StatementTEST StatementUNITS StatementWEIGHT Statement Missing ValuesModel SpecificationModel FittingSurvey Design InformationLogistic Regression Models and ParametersVariance EstimationDomain AnalysisHypothesis Testing and EstimationLinear Predictor, Predicted Probability, and Confidence LimitsOutput Data SetsDisplayed OutputODS Table NamesODS Graphics
Missing ValuesModel SpecificationModel FittingSurvey Design InformationLogistic Regression Models and ParametersVariance EstimationDomain AnalysisHypothesis Testing and EstimationLinear Predictor, Predicted Probability, and Confidence LimitsOutput Data SetsDisplayed OutputODS Table NamesODS Graphics