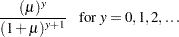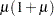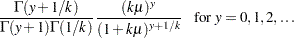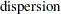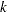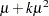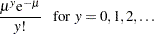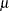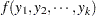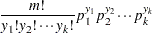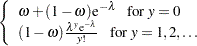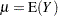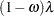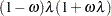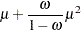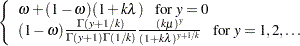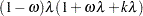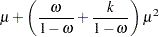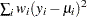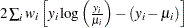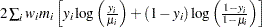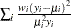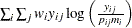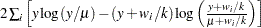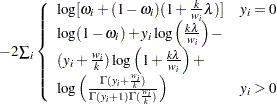The GENMOD Procedure
-
Overview

-
Getting Started

-
Syntax
 PROC GENMOD Statement ASSESS Statement BAYES Statement BY Statement CLASS Statement CONTRAST Statement DEVIANCE Statement EFFECTPLOT Statement ESTIMATE Statement EXACT Statement EXACTOPTIONS Statement FREQ Statement FWDLINK Statement INVLINK Statement LSMEANS Statement LSMESTIMATE Statement MODEL Statement OUTPUT Statement Programming Statements REPEATED Statement SLICE Statement STORE Statement STRATA Statement VARIANCE Statement WEIGHT Statement ZEROMODEL Statement
PROC GENMOD Statement ASSESS Statement BAYES Statement BY Statement CLASS Statement CONTRAST Statement DEVIANCE Statement EFFECTPLOT Statement ESTIMATE Statement EXACT Statement EXACTOPTIONS Statement FREQ Statement FWDLINK Statement INVLINK Statement LSMEANS Statement LSMESTIMATE Statement MODEL Statement OUTPUT Statement Programming Statements REPEATED Statement SLICE Statement STORE Statement STRATA Statement VARIANCE Statement WEIGHT Statement ZEROMODEL Statement -
Details
 Generalized Linear Models Theory Specification of Effects Parameterization Used in PROC GENMOD Type 1 Analysis Type 3 Analysis Confidence Intervals for Parameters F Statistics Lagrange Multiplier Statistics Predicted Values of the Mean Residuals Multinomial Models Zero-Inflated Models Generalized Estimating Equations Assessment of Models Based on Aggregates of Residuals Case Deletion Diagnostic Statistics Bayesian Analysis Exact Logistic and Exact Poisson Regression Missing Values Displayed Output for Classical Analysis Displayed Output for Bayesian Analysis Displayed Output for Exact Analysis ODS Table Names ODS Graphics
Generalized Linear Models Theory Specification of Effects Parameterization Used in PROC GENMOD Type 1 Analysis Type 3 Analysis Confidence Intervals for Parameters F Statistics Lagrange Multiplier Statistics Predicted Values of the Mean Residuals Multinomial Models Zero-Inflated Models Generalized Estimating Equations Assessment of Models Based on Aggregates of Residuals Case Deletion Diagnostic Statistics Bayesian Analysis Exact Logistic and Exact Poisson Regression Missing Values Displayed Output for Classical Analysis Displayed Output for Bayesian Analysis Displayed Output for Exact Analysis ODS Table Names ODS Graphics -
Examples
 Logistic Regression Normal Regression, Log Link Gamma Distribution Applied to Life Data Ordinal Model for Multinomial Data GEE for Binary Data with Logit Link Function Log Odds Ratios and the ALR Algorithm Log-Linear Model for Count Data Model Assessment of Multiple Regression Using Aggregates of Residuals Assessment of a Marginal Model for Dependent Data Bayesian Analysis of a Poisson Regression Model Exact Poisson Regression
Logistic Regression Normal Regression, Log Link Gamma Distribution Applied to Life Data Ordinal Model for Multinomial Data GEE for Binary Data with Logit Link Function Log Odds Ratios and the ALR Algorithm Log-Linear Model for Count Data Model Assessment of Multiple Regression Using Aggregates of Residuals Assessment of a Marginal Model for Dependent Data Bayesian Analysis of a Poisson Regression Model Exact Poisson Regression - References
| Generalized Linear Models Theory |
This is a brief introduction to the theory of generalized linear models.
Response Probability Distributions
In generalized linear models, the response is assumed to possess a probability distribution of the exponential form. That is, the probability density of the response 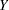 for continuous response variables, or the probability function for discrete responses, can be expressed as
for continuous response variables, or the probability function for discrete responses, can be expressed as
 |
for some functions  ,
, 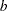 , and
, and  that determine the specific distribution. For fixed
that determine the specific distribution. For fixed  , this is a one-parameter exponential family of distributions. The functions
, this is a one-parameter exponential family of distributions. The functions  and
and  are such that
are such that  and
and 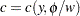 , where
, where 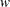 is a known weight for each observation. A variable representing
is a known weight for each observation. A variable representing 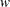 in the input data set can be specified in the WEIGHT statement. If no WEIGHT statement is specified,
in the input data set can be specified in the WEIGHT statement. If no WEIGHT statement is specified,  for all observations.
for all observations.
Standard theory for this type of distribution gives expressions for the mean and variance of 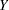 :
:
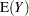 |
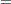 |
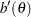 |
|||
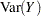 |
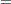 |
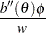 |
where the primes denote derivatives with respect to 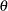 . If
. If 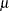 represents the mean of
represents the mean of 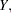 then the variance expressed as a function of the mean is
then the variance expressed as a function of the mean is
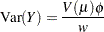 |
where  is the variance function.
is the variance function.
Probability distributions of the response 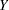 in generalized linear models are usually parameterized in terms of the mean
in generalized linear models are usually parameterized in terms of the mean 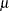 and dispersion parameter
and dispersion parameter  instead of the natural parameter
instead of the natural parameter 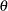 . The probability distributions that are available in the GENMOD procedure are shown in the following list. The zero-inflated Poisson and zero-inflated negative binomial distributions are not generalized linear models. However, the zero-inflated distributions are included in PROC GENMOD since they are useful extensions of generalized linear models. See Long (1997) for a discussion of the zero-inflated Poisson and zero-inflated negative binomial distributions. The PROC GENMOD scale parameter and the variance of
. The probability distributions that are available in the GENMOD procedure are shown in the following list. The zero-inflated Poisson and zero-inflated negative binomial distributions are not generalized linear models. However, the zero-inflated distributions are included in PROC GENMOD since they are useful extensions of generalized linear models. See Long (1997) for a discussion of the zero-inflated Poisson and zero-inflated negative binomial distributions. The PROC GENMOD scale parameter and the variance of 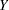 are also shown.
are also shown.
The negative binomial and the zero-inflated negative binomial distributions contain a parameter 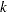 , called the negative binomial dispersion parameter. This is not the same as the generalized linear model dispersion
, called the negative binomial dispersion parameter. This is not the same as the generalized linear model dispersion  , but it is an additional distribution parameter that must be estimated or set to a fixed value.
, but it is an additional distribution parameter that must be estimated or set to a fixed value.
For the binomial distribution, the response is the binomial proportion 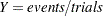 . The variance function is
. The variance function is  , and the binomial trials parameter
, and the binomial trials parameter  is regarded as a weight
is regarded as a weight 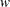 .
.
If a weight variable is present,  is replaced with
is replaced with 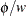 , where
, where 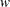 is the weight variable.
is the weight variable.
PROC GENMOD works with a scale parameter that is related to the exponential family dispersion parameter  instead of working with
instead of working with  itself. The scale parameters are related to the dispersion parameter as shown previously with the probability distribution definitions. Thus, the scale parameter output in the "Analysis of Parameter Estimates" table is related to the exponential family dispersion parameter. If you specify a constant scale parameter with the SCALE= option in the MODEL statement, it is also related to the exponential family dispersion parameter in the same way.
itself. The scale parameters are related to the dispersion parameter as shown previously with the probability distribution definitions. Thus, the scale parameter output in the "Analysis of Parameter Estimates" table is related to the exponential family dispersion parameter. If you specify a constant scale parameter with the SCALE= option in the MODEL statement, it is also related to the exponential family dispersion parameter in the same way.
Link Function
For distributions other than the zero-inflated Poisson or zero-inflated negative binomial, the mean 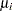 of the response in the
of the response in the 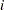 th observation is related to a linear predictor through a monotonic differentiable link function
th observation is related to a linear predictor through a monotonic differentiable link function 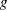 .
.
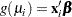 |
Here, 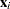 is a fixed known vector of explanatory variables, and
is a fixed known vector of explanatory variables, and 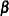 is a vector of unknown parameters.
is a vector of unknown parameters.
There are two link functions and linear predictors associated with zero-inflated distributions: one for the zero inflation probability 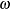 , and another for the mean parameter
, and another for the mean parameter 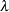 . See the section Zero-Inflated Models for more details about zero-inflated distributions.
. See the section Zero-Inflated Models for more details about zero-inflated distributions.
Log-Likelihood Functions
Log-likelihood functions for the distributions that are available in the procedure are parameterized in terms of the means 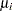 and the dispersion parameter
and the dispersion parameter  . Zero-inflated log likelihoods are parameterized in terms two parameters,
. Zero-inflated log likelihoods are parameterized in terms two parameters, 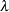 and
and 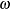 . The parameter
. The parameter 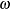 is the zero-inflation probability, and
is the zero-inflation probability, and 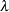 is a function of the distribution mean. The relationship between the mean of the zero-inflated Poisson and zero-inflated negative binomial distributions and the parameter
is a function of the distribution mean. The relationship between the mean of the zero-inflated Poisson and zero-inflated negative binomial distributions and the parameter 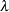 is defined in the section Response Probability Distributions. The term
is defined in the section Response Probability Distributions. The term  represents the response for the
represents the response for the 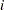 th observation, and
th observation, and 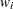 represents the known dispersion weight. The log-likelihood functions are of the form
represents the known dispersion weight. The log-likelihood functions are of the form
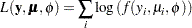 |
where the sum is over the observations. The forms of the individual contributions
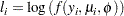 |
are shown in the following list; the parameterizations are expressed in terms of the mean and dispersion parameters.
For the discrete distributions (binomial, multinomial, negative binomial, and Poisson), the functions computed as the sum of the 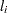 terms are not proper log-likelihood functions, since terms involving binomial coefficients or factorials of the observed counts are dropped from the computation of the log likelihood, and a dispersion parameter
terms are not proper log-likelihood functions, since terms involving binomial coefficients or factorials of the observed counts are dropped from the computation of the log likelihood, and a dispersion parameter  is included in the computation. Deletion of factorial terms and inclusion of a dispersion parameter do not affect parameter estimates or their estimated covariances for these distributions, and this is the function used in maximum likelihood estimation. The value of
is included in the computation. Deletion of factorial terms and inclusion of a dispersion parameter do not affect parameter estimates or their estimated covariances for these distributions, and this is the function used in maximum likelihood estimation. The value of  used in computing the reported log-likelihood function is either the final estimated value, or the fixed value, if the dispersion parameter is fixed. Even though it is not a proper log-likelihood function in all cases, the function computed as the sum of the
used in computing the reported log-likelihood function is either the final estimated value, or the fixed value, if the dispersion parameter is fixed. Even though it is not a proper log-likelihood function in all cases, the function computed as the sum of the 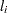 terms is reported in the output as the log likelihood. The proper log-likelihood function is also computed as the sum of the
terms is reported in the output as the log likelihood. The proper log-likelihood function is also computed as the sum of the 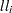 terms in the following list, and it is reported as the full log likelihood in the output.
terms in the following list, and it is reported as the full log likelihood in the output.
-
Normal:
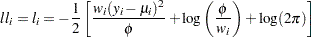
-
Inverse Gaussian:
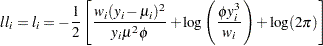
-
Gamma:
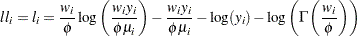
-
Negative binomial:
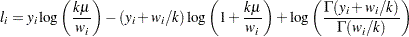
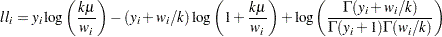
-
Poisson:
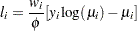
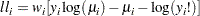
-
Binomial:
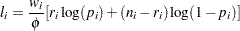
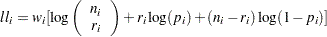
-
Multinomial (k categories):
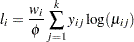
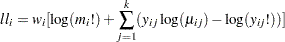
-
Zero-inflated Poisson:
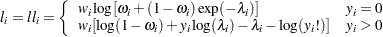
-
Zero-inflated negative binomial:

Maximum Likelihood Fitting
The GENMOD procedure uses a ridge-stabilized Newton-Raphson algorithm to maximize the log-likelihood function 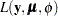 with respect to the regression parameters. By default, the procedure also produces maximum likelihood estimates of the scale parameter as defined in the section Response Probability Distributions for the normal, inverse Gaussian, negative binomial, and gamma distributions.
with respect to the regression parameters. By default, the procedure also produces maximum likelihood estimates of the scale parameter as defined in the section Response Probability Distributions for the normal, inverse Gaussian, negative binomial, and gamma distributions.
On the  th iteration, the algorithm updates the parameter vector
th iteration, the algorithm updates the parameter vector 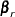 with
with
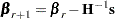 |
where 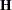 is the Hessian (second derivative) matrix, and
is the Hessian (second derivative) matrix, and  is the gradient (first derivative) vector of the log-likelihood function, both evaluated at the current value of the parameter vector. That is,
is the gradient (first derivative) vector of the log-likelihood function, both evaluated at the current value of the parameter vector. That is,
 |
and
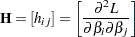 |
In some cases, the scale parameter is estimated by maximum likelihood. In these cases, elements corresponding to the scale parameter are computed and included in  and
and 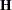 .
.
If 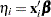 is the linear predictor for observation
is the linear predictor for observation 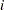 and
and 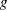 is the link function, then
is the link function, then 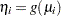 , so that
, so that 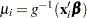 is an estimate of the mean of the
is an estimate of the mean of the 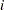 th observation, obtained from an estimate of the parameter vector
th observation, obtained from an estimate of the parameter vector 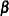 .
.
The gradient vector and Hessian matrix for the regression parameters are given by
 |
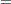 |
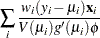 |
|||
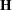 |
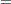 |
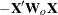 |
where 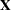 is the design matrix,
is the design matrix, 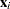 is the transpose of the
is the transpose of the 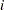 th row of X, and
th row of X, and  is the variance function. The matrix
is the variance function. The matrix 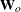 is diagonal with its
is diagonal with its 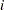 th diagonal element
th diagonal element
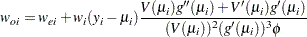 |
where
 |
The primes denote derivatives of 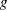 and
and  with respect to
with respect to 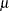 . The negative of
. The negative of 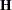 is called the observed information matrix. The expected value of
is called the observed information matrix. The expected value of 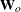 is a diagonal matrix
is a diagonal matrix  with diagonal values
with diagonal values 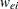 . If you replace
. If you replace 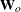 with
with  , then the negative of
, then the negative of 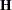 is called the expected information matrix.
is called the expected information matrix.  is the weight matrix for the Fisher scoring method of fitting. Either
is the weight matrix for the Fisher scoring method of fitting. Either 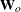 or
or  can be used in the update equation. The GENMOD procedure uses Fisher scoring for iterations up to the number specified by the SCORING option in the MODEL statement, and it uses the observed information matrix on additional iterations.
can be used in the update equation. The GENMOD procedure uses Fisher scoring for iterations up to the number specified by the SCORING option in the MODEL statement, and it uses the observed information matrix on additional iterations.
Covariance and Correlation Matrix
The estimated covariance matrix of the parameter estimator is given by
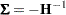 |
where 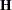 is the Hessian matrix evaluated using the parameter estimates on the last iteration. Note that the dispersion parameter, whether estimated or specified, is incorporated into
is the Hessian matrix evaluated using the parameter estimates on the last iteration. Note that the dispersion parameter, whether estimated or specified, is incorporated into 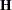 . Rows and columns corresponding to aliased parameters are not included in
. Rows and columns corresponding to aliased parameters are not included in 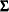 .
.
The correlation matrix is the normalized covariance matrix. That is, if 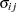 is an element of
is an element of 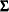 , then the corresponding element of the correlation matrix is
, then the corresponding element of the correlation matrix is 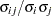 , where
, where 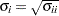 .
.
Goodness of Fit
Two statistics that are helpful in assessing the goodness of fit of a given generalized linear model are the scaled deviance and Pearson’s chi-square statistic. For a fixed value of the dispersion parameter  , the scaled deviance is defined to be twice the difference between the maximum achievable log likelihood and the log likelihood at the maximum likelihood estimates of the regression parameters.
, the scaled deviance is defined to be twice the difference between the maximum achievable log likelihood and the log likelihood at the maximum likelihood estimates of the regression parameters.
Note that these statistics are not valid for GEE models.
If 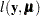 is the log-likelihood function expressed as a function of the predicted mean values
is the log-likelihood function expressed as a function of the predicted mean values 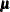 and the vector
and the vector 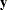 of response values, then the scaled deviance is defined by
of response values, then the scaled deviance is defined by
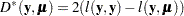 |
For specific distributions, this can be expressed as
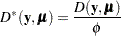 |
where  is the deviance. The following table displays the deviance for each of the probability distributions available in PROC GENMOD. The deviance cannot be directly calculated for zero-inflated models. Twice the negative of the log likelihood is reported instead of the proper deviance for the zero-inflated Poisson and zero-inflated negative binomial.
is the deviance. The following table displays the deviance for each of the probability distributions available in PROC GENMOD. The deviance cannot be directly calculated for zero-inflated models. Twice the negative of the log likelihood is reported instead of the proper deviance for the zero-inflated Poisson and zero-inflated negative binomial.
Distribution |
Deviance |
|---|---|
Normal |
|
Poisson |
|
Binomial |
|
Gamma |
|
Inverse Gaussian |
|
Multinomial |
|
Negative binomial |
|
Zero-inflated Poisson |
|
Zero-inflated negative binomial |
|
In the binomial case, 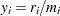 , where
, where  is a binomial count and
is a binomial count and  is the binomial number of trials parameter.
is the binomial number of trials parameter.
In the multinomial case, 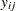 refers to the observed number of occurrences of the
refers to the observed number of occurrences of the 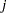 th category for the
th category for the 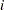 th subpopulation defined by the AGGREGATE= variable,
th subpopulation defined by the AGGREGATE= variable,  is the total number in the
is the total number in the 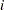 th subpopulation, and
th subpopulation, and 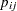 is the category probability.
is the category probability.
Pearson’s chi-square statistic is defined as
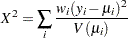 |
and the scaled Pearson’s chi-square is  .
.
The scaled version of both of these statistics, under certain regularity conditions, has a limiting chi-square distribution, with degrees of freedom equal to the number of observations minus the number of parameters estimated. The scaled version can be used as an approximate guide to the goodness of fit of a given model. Use caution before applying these statistics to ensure that all the conditions for the asymptotic distributions hold. McCullagh and Nelder (1989) advise that differences in deviances for nested models can be better approximated by chi-square distributions than the deviances can themselves.
In cases where the dispersion parameter is not known, an estimate can be used to obtain an approximation to the scaled deviance and Pearson’s chi-square statistic. One strategy is to fit a model that contains a sufficient number of parameters so that all systematic variation is removed, estimate  from this model, and then use this estimate in computing the scaled deviance of submodels. The deviance or Pearson’s chi-square divided by its degrees of freedom is sometimes used as an estimate of the dispersion parameter
from this model, and then use this estimate in computing the scaled deviance of submodels. The deviance or Pearson’s chi-square divided by its degrees of freedom is sometimes used as an estimate of the dispersion parameter  . For example, since the limiting chi-square distribution of the scaled deviance
. For example, since the limiting chi-square distribution of the scaled deviance 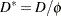 has
has 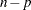 degrees of freedom, where
degrees of freedom, where  is the number of observations and
is the number of observations and 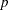 is the number of parameters, equating
is the number of parameters, equating 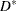 to its mean and solving for
to its mean and solving for  yields
yields 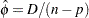 . Similarly, an estimate of
. Similarly, an estimate of  based on Pearson’s chi-square
based on Pearson’s chi-square 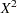 is
is 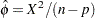 . Alternatively, a maximum likelihood estimate of
. Alternatively, a maximum likelihood estimate of  can be computed by the procedure, if desired. See the discussion in the section Type 1 Analysis for more about the estimation of the dispersion parameter.
can be computed by the procedure, if desired. See the discussion in the section Type 1 Analysis for more about the estimation of the dispersion parameter.
Other Fit Statistics
The Akaike information criterion (AIC) is a measure of goodness of model fit that balances model fit against model simplicity. AIC has the form
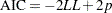 |
where 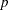 is the number of parameters estimated in the model, and
is the number of parameters estimated in the model, and 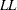 is the log likelihood evaluated at the value of the estimated parameters. An alternative form is the corrected AIC given by
is the log likelihood evaluated at the value of the estimated parameters. An alternative form is the corrected AIC given by
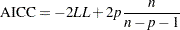 |
where  is the total number of observations used.
is the total number of observations used.
The Bayesian information criterion (BIC) is a similar measure. BIC is defined by
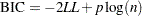 |
See Akaike (1981, 1979) for details of AIC and BIC. See Simonoff (2003) for a discussion of using AIC, AICC, and BIC with generalized linear models. These criteria are useful in selecting among regression models, with smaller values representing better model fit. PROC GENMOD uses the full log likelihoods defined in the section Log-Likelihood Functions, with all terms included, for computing all of the criteria.
Dispersion Parameter
There are several options available in PROC GENMOD for handling the exponential distribution dispersion parameter. The NOSCALE and SCALE options in the MODEL statement affect the way in which the dispersion parameter is treated. If you specify the SCALE=DEVIANCE option, the dispersion parameter is estimated by the deviance divided by its degrees of freedom. If you specify the SCALE=PEARSON option, the dispersion parameter is estimated by Pearson’s chi-square statistic divided by its degrees of freedom.
Otherwise, values of the SCALE and NOSCALE options and the resultant actions are displayed in the following table.
NOSCALE |
SCALE=value |
Action |
|---|---|---|
Present |
Present |
Scale fixed at value |
Present |
Not present |
Scale fixed at 1 |
Not present |
Not present |
Scale estimated by ML |
Not present |
Present |
Scale estimated by ML, |
starting point at value |
||
Present (negative binomial) |
Not present |
|
The meaning of the scale parameter displayed in the "Analysis Of Parameter Estimates" table is different for the gamma distribution than for the other distributions. The relation of the scale parameter as used by PROC GENMOD to the exponential family dispersion parameter  is displayed in the following table. For the binomial and Poisson distributions,
is displayed in the following table. For the binomial and Poisson distributions,  is the overdispersion parameter, as defined in the "Overdispersion" section, which follows.
is the overdispersion parameter, as defined in the "Overdispersion" section, which follows.
Distribution |
Scale |
|---|---|
Normal |
|
Inverse Gaussian |
|
Gamma |
|
Binomial |
|
Poisson |
|
In the case of the negative binomial distribution, PROC GENMOD reports the "dispersion" parameter estimated by maximum likelihood. This is the negative binomial parameter 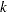 defined in the section Response Probability Distributions.
defined in the section Response Probability Distributions.
Overdispersion
Overdispersion is a phenomenon that sometimes occurs in data that are modeled with the binomial or Poisson distributions. If the estimate of dispersion after fitting, as measured by the deviance or Pearson’s chi-square, divided by the degrees of freedom, is not near 1, then the data might be overdispersed if the dispersion estimate is greater than 1 or underdispersed if the dispersion estimate is less than 1. A simple way to model this situation is to allow the variance functions of these distributions to have a multiplicative overdispersion factor  :
:
Binomial:

Poisson:
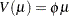
An alternative method to allow for overdispersion in the Poisson distribution is to fit a negative binomial distribution, where 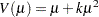 , instead of the Poisson. The parameter
, instead of the Poisson. The parameter 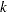 can be estimated by maximum likelihood, thus allowing for overdispersion of a specific form. This is different from the multiplicative overdispersion factor
can be estimated by maximum likelihood, thus allowing for overdispersion of a specific form. This is different from the multiplicative overdispersion factor  , which can accommodate many forms of overdispersion.
, which can accommodate many forms of overdispersion.
The models are fit in the usual way, and the parameter estimates are not affected by the value of  . The covariance matrix, however, is multiplied by
. The covariance matrix, however, is multiplied by  , and the scaled deviance and log likelihoods used in likelihood ratio tests are divided by
, and the scaled deviance and log likelihoods used in likelihood ratio tests are divided by  . The profile likelihood function used in computing confidence intervals is also divided by
. The profile likelihood function used in computing confidence intervals is also divided by  . If you specify a WEIGHT statement,
. If you specify a WEIGHT statement,  is divided by the value of the WEIGHT variable for each observation. This has the effect of multiplying the contributions of the log-likelihood function, the gradient, and the Hessian by the value of the WEIGHT variable for each observation.
is divided by the value of the WEIGHT variable for each observation. This has the effect of multiplying the contributions of the log-likelihood function, the gradient, and the Hessian by the value of the WEIGHT variable for each observation.
The SCALE= option in the MODEL statement enables you to specify a value of 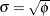 for the binomial and Poisson distributions. If you specify the SCALE=DEVIANCE option in the MODEL statement, the procedure uses the deviance divided by degrees of freedom as an estimate of
for the binomial and Poisson distributions. If you specify the SCALE=DEVIANCE option in the MODEL statement, the procedure uses the deviance divided by degrees of freedom as an estimate of  , and all statistics are adjusted appropriately. You can use Pearson’s chi-square instead of the deviance by specifying the SCALE=PEARSON option.
, and all statistics are adjusted appropriately. You can use Pearson’s chi-square instead of the deviance by specifying the SCALE=PEARSON option.
The function obtained by dividing a log-likelihood function for the binomial or Poisson distribution by a dispersion parameter is not a legitimate log-likelihood function. It is an example of a quasi-likelihood function. Most of the asymptotic theory for log likelihoods also applies to quasi-likelihoods, which justifies computing standard errors and likelihood ratio statistics by using quasi-likelihoods instead of proper log likelihoods. See McCullagh and Nelder (1989, Chapter 9), McCullagh (1983), and Hardin and Hilbe (2003) for details on quasi-likelihood functions.
Although the estimate of the dispersion parameter is often used to indicate overdispersion or underdispersion, this estimate might also indicate other problems such as an incorrectly specified model or outliers in the data. You should carefully assess whether this type of model is appropriate for your data.
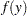
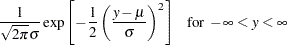
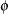
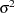


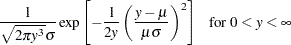
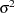
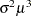



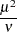
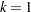 .
.