The NLMIXED Procedure
-
Overview

-
Getting Started

-
Syntax

-
Details
 Modeling Assumptions and NotationIntegral ApproximationsBuilt-in Log-Likelihood FunctionsHierarchical Model SpecificationOptimization AlgorithmsFinite-Difference Approximations of DerivativesHessian ScalingActive Set MethodsLine-Search MethodsRestricting the Step LengthComputational ProblemsCovariance MatrixPredictionComputational ResourcesDisplayed OutputODS Table Names
Modeling Assumptions and NotationIntegral ApproximationsBuilt-in Log-Likelihood FunctionsHierarchical Model SpecificationOptimization AlgorithmsFinite-Difference Approximations of DerivativesHessian ScalingActive Set MethodsLine-Search MethodsRestricting the Step LengthComputational ProblemsCovariance MatrixPredictionComputational ResourcesDisplayed OutputODS Table Names -
Examples

- References
Covariance Matrix
The estimated covariance matrix of the parameter estimates is computed as the inverse Hessian matrix, and for unconstrained
problems it should be positive definite. If the final parameter estimates are subjected to 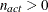 active linear inequality constraints, the formulas of the covariance matrices are modified similar to Gallant (1987) and Cramer (1986, p. 38) and additionally generalized for applications with singular matrices.
active linear inequality constraints, the formulas of the covariance matrices are modified similar to Gallant (1987) and Cramer (1986, p. 38) and additionally generalized for applications with singular matrices.
There are several steps available that enable you to tune the rank calculations of the covariance matrix.
-
You can use the ASINGULAR= , MSINGULAR= , and VSINGULAR= options to set three singularity criteria for the inversion of the Hessian matrix
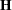 . The singularity criterion used for the inversion is
. The singularity criterion used for the inversion is
![\[ |d_{j,j}| \le \max (\mbox{ASING}, \mbox{VSING} * |H_{j,j}|, \mbox{MSING} * \max (|H_{1,1}|,\ldots ,|H_{n,n}|)) \]](images/statug_nlmixed0300.png)
where
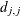 is the diagonal pivot of the matrix
is the diagonal pivot of the matrix 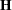 , and ASING, VSING, and MSING are the specified values of the ASINGULAR=
, VSINGULAR=
, and MSINGULAR=
options, respectively. The default values are as follows:
, and ASING, VSING, and MSING are the specified values of the ASINGULAR=
, VSINGULAR=
, and MSINGULAR=
options, respectively. The default values are as follows:
Note that, in many cases, a normalized matrix
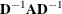 is decomposed, and the singularity criteria are modified correspondingly.
is decomposed, and the singularity criteria are modified correspondingly.
-
If the matrix
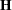 is found to be singular in the first step, a generalized inverse is computed. Depending on the G4=
option, either a generalized inverse satisfying all four Moore-Penrose conditions is computed (a
is found to be singular in the first step, a generalized inverse is computed. Depending on the G4=
option, either a generalized inverse satisfying all four Moore-Penrose conditions is computed (a 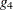 -inverse) or a generalized inverse satisfying only two Moore-Penrose conditions is computed (a
-inverse) or a generalized inverse satisfying only two Moore-Penrose conditions is computed (a 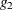 -inverse, Pringle and Rayner 1971). If the number of parameters n of the application is less than or equal to G4=
i, a
-inverse, Pringle and Rayner 1971). If the number of parameters n of the application is less than or equal to G4=
i, a 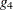 -inverse is computed; otherwise, only a
-inverse is computed; otherwise, only a 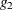 -inverse is computed. The
-inverse is computed. The 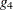 -inverse is computed by the (computationally very expensive but numerically stable) eigenvalue decomposition, and the
-inverse is computed by the (computationally very expensive but numerically stable) eigenvalue decomposition, and the 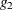 -inverse is computed by Gauss transformation. The
-inverse is computed by Gauss transformation. The 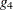 -inverse is computed using the eigenvalue decomposition
-inverse is computed using the eigenvalue decomposition 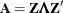 , where
, where 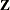 is the orthogonal matrix of eigenvectors and
is the orthogonal matrix of eigenvectors and 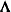 is the diagonal matrix of eigenvalues,
is the diagonal matrix of eigenvalues, 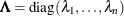 . The
. The 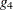 -inverse of
-inverse of 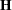 is set to
is set to
![\[ \mb{A}^- = \mb{Z} \bLambda ^- \mb{Z}^\prime \]](images/statug_nlmixed0309.png)
where the diagonal matrix
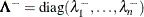 is defined using the COVSING=
option:
is defined using the COVSING=
option:
![\[ \lambda ^-_ i = \left\{ \begin{array}{ll} 1 / \lambda _ i & \mr{if} \, \, |\lambda _ i| > \mr{COVSING} \\ 0 & \mr{if} \, \, |\lambda _ i| \le \mr{COVSING} \end{array} \right. \]](images/statug_nlmixed0311.png)
If you do not specify the COVSING= option, the nr smallest eigenvalues are set to zero, where nr is the number of rank deficiencies found in the first step.
For optimization techniques that do not use second-order derivatives, the covariance matrix is computed using finite-difference approximations of the derivatives.