The VARMAX Procedure
- Overview
-
Getting Started

-
Syntax

-
Details
 Missing ValuesVARMAX ModelDynamic Simultaneous Equations ModelingImpulse Response FunctionForecastingTentative Order SelectionVAR and VARX ModelingBayesian VAR and VARX ModelingVARMA and VARMAX ModelingModel Diagnostic ChecksCointegrationVector Error Correction ModelingI(2) ModelMultivariate GARCH ModelingOutput Data SetsOUT= Data SetOUTEST= Data SetOUTHT= Data SetOUTSTAT= Data SetPrinted OutputODS Table NamesODS GraphicsComputational Issues
Missing ValuesVARMAX ModelDynamic Simultaneous Equations ModelingImpulse Response FunctionForecastingTentative Order SelectionVAR and VARX ModelingBayesian VAR and VARX ModelingVARMA and VARMAX ModelingModel Diagnostic ChecksCointegrationVector Error Correction ModelingI(2) ModelMultivariate GARCH ModelingOutput Data SetsOUT= Data SetOUTEST= Data SetOUTHT= Data SetOUTSTAT= Data SetPrinted OutputODS Table NamesODS GraphicsComputational Issues -
Examples

- References
Vector Autoregressive Process
Let ![]()
![]() denote a
denote a ![]() -dimensional time series vector of random variables of interest. The
-dimensional time series vector of random variables of interest. The ![]() th-order VAR process is written as
th-order VAR process is written as
|
|
where the ![]() is a vector white noise process with
is a vector white noise process with ![]() such that
such that ![]() ,
, ![]() , and
, and ![]() for
for ![]() ;
; ![]() is a constant vector and
is a constant vector and ![]() is a
is a ![]() matrix.
matrix.
Analyzing and modeling the series jointly enables you to understand the dynamic relationships over time among the series and to improve the accuracy of forecasts for individual series by using the additional information available from the related series and their forecasts.
Example of Vector Autoregressive Model
Consider the first-order stationary bivariate vector autoregressive model
|
|
The following IML procedure statements simulate a bivariate vector time series from this model to provide test data for the VARMAX procedure:
proc iml;
sig = {1.0 0.5, 0.5 1.25};
phi = {1.2 -0.5, 0.6 0.3};
/* simulate the vector time series */
call varmasim(y,phi) sigma = sig n = 100 seed = 34657;
cn = {'y1' 'y2'};
create simul1 from y[colname=cn];
append from y;
quit;
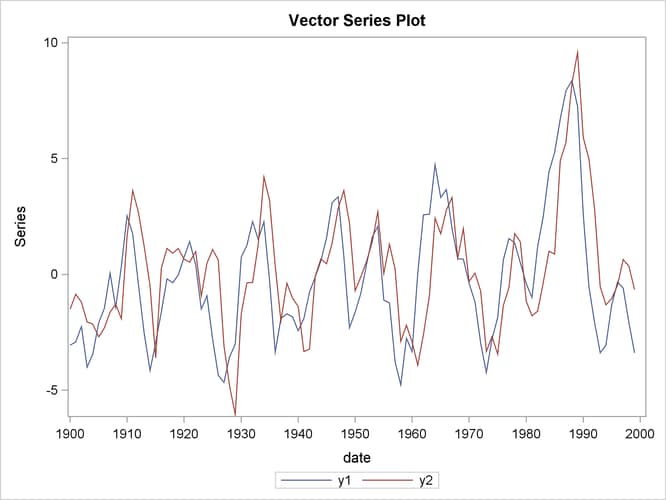
The following statements plot the simulated vector time series ![]() shown in Figure 36.1:
shown in Figure 36.1:
data simul1; set simul1; date = intnx( 'year', '01jan1900'd, _n_-1 ); format date year4.; run; ods graphics on; proc timeseries data=simul1 vectorplot=series; id date interval=year; var y1 y2; run;
Figure 36.1: Plot of Generated Data Process
The following statements fit a VAR(1) model to the simulated data. First, you specify the input data set in the PROC VARMAX
statement. Then, you use the MODEL statement to designate the dependent variables, ![]() and
and ![]() . To estimate a VAR model with mean zero, you specify the order of the autoregressive model with the P= option and the NOINT
option. The MODEL statement fits the model to the data and prints parameter estimates and their significance. The PRINT=ESTIMATES
option prints the matrix form of parameter estimates, and the PRINT=DIAGNOSE option prints various diagnostic tests. The LAGMAX=3
option is used to print the output for the residual diagnostic checks.
. To estimate a VAR model with mean zero, you specify the order of the autoregressive model with the P= option and the NOINT
option. The MODEL statement fits the model to the data and prints parameter estimates and their significance. The PRINT=ESTIMATES
option prints the matrix form of parameter estimates, and the PRINT=DIAGNOSE option prints various diagnostic tests. The LAGMAX=3
option is used to print the output for the residual diagnostic checks.
To output the forecasts to a data set, you specify the OUTPUT statement with the OUT= option. If you want to forecast five steps ahead, you use the LEAD=5 option. The ID statement specifies the yearly interval between observations and provides the Time column in the forecast output.
The VARMAX procedure output is shown in Figure 36.2 through Figure 36.10.
/*--- Vector Autoregressive Model ---*/
proc varmax data=simul1;
id date interval=year;
model y1 y2 / p=1 noint lagmax=3
print=(estimates diagnose);
output out=for lead=5;
run;
Figure 36.2: Descriptive Statistics
| Number of Observations | 100 |
|---|---|
| Number of Pairwise Missing | 0 |
| Simple Summary Statistics | ||||||
|---|---|---|---|---|---|---|
| Variable | Type | N | Mean | Standard Deviation |
Min | Max |
| y1 | Dependent | 100 | -0.21653 | 2.78210 | -4.75826 | 8.37032 |
| y2 | Dependent | 100 | 0.16905 | 2.58184 | -6.04718 | 9.58487 |
The VARMAX procedure first displays descriptive statistics. The Type column specifies that the variables are dependent variables. The column N stands for the number of nonmissing observations.
Figure 36.3 shows the type and the estimation method of the fitted model for the simulated data. It also shows the AR coefficient matrix in terms of lag 1, the parameter estimates, and their significance, which can indicate how well the model fits the data.
The second table schematically represents the parameter estimates and allows for easy verification of their significance in matrix form.
In the last table, the first column gives the left-hand-side variable of the equation; the second column is the parameter
name AR![]() , which indicates the (
, which indicates the (![]() )th element of the lag
)th element of the lag ![]() autoregressive coefficient; the last column is the regressor that corresponds to the displayed parameter.
autoregressive coefficient; the last column is the regressor that corresponds to the displayed parameter.
Figure 36.3: Model Type and Parameter Estimates
| Type of Model | VAR(1) |
|---|---|
| Estimation Method | Least Squares Estimation |
| AR | |||
|---|---|---|---|
| Lag | Variable | y1 | y2 |
| 1 | y1 | 1.15977 | -0.51058 |
| y2 | 0.54634 | 0.38499 | |
| Schematic Representation |
|
|---|---|
| Variable/Lag | AR1 |
| y1 | +- |
| y2 | ++ |
| + is > 2*std error, - is < -2*std error, . is between, * is N/A | |
| Model Parameter Estimates | ||||||
|---|---|---|---|---|---|---|
| Equation | Parameter | Estimate | Standard Error |
t Value | Pr > |t| | Variable |
| y1 | AR1_1_1 | 1.15977 | 0.05508 | 21.06 | 0.0001 | y1(t-1) |
| AR1_1_2 | -0.51058 | 0.05898 | -8.66 | 0.0001 | y2(t-1) | |
| y2 | AR1_2_1 | 0.54634 | 0.05779 | 9.45 | 0.0001 | y1(t-1) |
| AR1_2_2 | 0.38499 | 0.06188 | 6.22 | 0.0001 | y2(t-1) | |
The fitted VAR(1) model with estimated standard errors in parentheses is given as
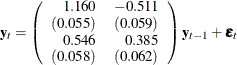 |
Clearly, all parameter estimates in the coefficient matrix ![]() are significant.
are significant.
The model can also be written as two univariate regression equations.
|
|
|
|
|
|
|
|
The table in Figure 36.4 shows the innovation covariance matrix estimates and the various information criteria results. The smaller value of information
criteria fits the data better when it is compared to other models. The variable names in the covariance matrix are printed
for convenience; ![]() means the innovation for
means the innovation for ![]() , and
, and ![]() means the innovation for
means the innovation for ![]() .
.
Figure 36.4: Innovation Covariance Estimates and Information Criteria
| Covariances of Innovations | ||
|---|---|---|
| Variable | y1 | y2 |
| y1 | 1.28875 | 0.39751 |
| y2 | 0.39751 | 1.41839 |
| Information Criteria | |
|---|---|
| AICC | 0.554443 |
| HQC | 0.595201 |
| AIC | 0.552777 |
| SBC | 0.65763 |
| FPEC | 1.738092 |
Figure 36.5 shows the cross covariances of the residuals. The values of the lag zero are slightly different from Figure 36.4 due to the different degrees of freedom.
Figure 36.5: Multivariate Diagnostic Checks
| Cross Covariances of Residuals | |||
|---|---|---|---|
| Lag | Variable | y1 | y2 |
| 0 | y1 | 1.26271 | 0.38948 |
| y2 | 0.38948 | 1.38974 | |
| 1 | y1 | 0.03121 | 0.05675 |
| y2 | -0.04646 | -0.05398 | |
| 2 | y1 | 0.08134 | 0.10599 |
| y2 | 0.03482 | -0.01549 | |
| 3 | y1 | 0.01644 | 0.11734 |
| y2 | 0.00609 | 0.11414 | |
Figure 36.6 and Figure 36.7 show tests for white noise residuals. The output shows that you cannot reject the null hypothesis that the residuals are uncorrelated.
Figure 36.6: Multivariate Diagnostic Checks Continued
| Cross Correlations of Residuals | |||
|---|---|---|---|
| Lag | Variable | y1 | y2 |
| 0 | y1 | 1.00000 | 0.29401 |
| y2 | 0.29401 | 1.00000 | |
| 1 | y1 | 0.02472 | 0.04284 |
| y2 | -0.03507 | -0.03884 | |
| 2 | y1 | 0.06442 | 0.08001 |
| y2 | 0.02628 | -0.01115 | |
| 3 | y1 | 0.01302 | 0.08858 |
| y2 | 0.00460 | 0.08213 | |
| Schematic Representation of Cross Correlations of Residuals |
||||
|---|---|---|---|---|
| Variable/Lag | 0 | 1 | 2 | 3 |
| y1 | ++ | .. | .. | .. |
| y2 | ++ | .. | .. | .. |
| + is > 2*std error, - is < -2*std error, . is between | ||||
Figure 36.7: Multivariate Diagnostic Checks Continued
| Portmanteau Test for Cross Correlations of Residuals |
|||
|---|---|---|---|
| Up To Lag | DF | Chi-Square | Pr > ChiSq |
| 2 | 4 | 1.58 | 0.8124 |
| 3 | 8 | 2.78 | 0.9473 |
The VARMAX procedure provides diagnostic checks for the univariate form of the equations. The table in Figure 36.8 describes how well each univariate equation fits the data. From two univariate regression equations in Figure 36.3, the values of ![]() in the second column are 0.84 and 0.80 for each equation. The standard deviations in the third column are the square roots
of the diagonal elements of the covariance matrix from Figure 36.4. The
in the second column are 0.84 and 0.80 for each equation. The standard deviations in the third column are the square roots
of the diagonal elements of the covariance matrix from Figure 36.4. The ![]() statistics are in the fourth column for hypotheses to test
statistics are in the fourth column for hypotheses to test ![]() and
and ![]() , respectively, where
, respectively, where ![]() is the
is the ![]() th element of the matrix
th element of the matrix ![]() . The last column shows the
. The last column shows the ![]() -values of the
-values of the ![]() statistics. The results show that each univariate model is significant.
statistics. The results show that each univariate model is significant.
Figure 36.8: Univariate Diagnostic Checks
| Univariate Model ANOVA Diagnostics | ||||
|---|---|---|---|---|
| Variable | R-Square | Standard Deviation |
F Value | Pr > F |
| y1 | 0.8351 | 1.13523 | 491.25 | <.0001 |
| y2 | 0.7906 | 1.19096 | 366.29 | <.0001 |
The check for white noise residuals in terms of the univariate equation is shown in Figure 36.9. This output contains information that indicates whether the residuals are correlated and heteroscedastic. In the first table,
the second column contains the Durbin-Watson test statistics to test the null hypothesis that the residuals are uncorrelated.
The third and fourth columns show the Jarque-Bera normality test statistics and their ![]() -values to test the null hypothesis that the residuals have normality. The last two columns show
-values to test the null hypothesis that the residuals have normality. The last two columns show ![]() statistics and their
statistics and their ![]() -values for ARCH(1) disturbances to test the null hypothesis that the residuals have equal covariances. The second table includes
-values for ARCH(1) disturbances to test the null hypothesis that the residuals have equal covariances. The second table includes
![]() statistics and their
statistics and their ![]() -values for AR(1), AR(1,2), AR(1,2,3) and AR(1,2,3,4) models of residuals to test the null hypothesis that the residuals are
uncorrelated.
-values for AR(1), AR(1,2), AR(1,2,3) and AR(1,2,3,4) models of residuals to test the null hypothesis that the residuals are
uncorrelated.
Figure 36.9: Univariate Diagnostic Checks Continued
| Univariate Model White Noise Diagnostics | |||||
|---|---|---|---|---|---|
| Variable | Durbin Watson |
Normality | ARCH | ||
| Chi-Square | Pr > ChiSq | F Value | Pr > F | ||
| y1 | 1.94534 | 3.56 | 0.1686 | 0.13 | 0.7199 |
| y2 | 2.06276 | 5.42 | 0.0667 | 2.10 | 0.1503 |
| Univariate Model AR Diagnostics | ||||||||
|---|---|---|---|---|---|---|---|---|
| Variable | AR1 | AR2 | AR3 | AR4 | ||||
| F Value | Pr > F | F Value | Pr > F | F Value | Pr > F | F Value | Pr > F | |
| y1 | 0.02 | 0.8980 | 0.14 | 0.8662 | 0.09 | 0.9629 | 0.82 | 0.5164 |
| y2 | 0.52 | 0.4709 | 0.41 | 0.6650 | 0.32 | 0.8136 | 0.32 | 0.8664 |
The table in Figure 36.10 gives forecasts, their prediction errors, and 95% confidence limits. See the section Forecasting for details.
Figure 36.10: Forecasts
| Forecasts | ||||||
|---|---|---|---|---|---|---|
| Variable | Obs | Time | Forecast | Standard Error |
95% Confidence Limits | |
| y1 | 101 | 2000 | -3.59212 | 1.13523 | -5.81713 | -1.36711 |
| 102 | 2001 | -3.09448 | 1.70915 | -6.44435 | 0.25539 | |
| 103 | 2002 | -2.17433 | 2.14472 | -6.37792 | 2.02925 | |
| 104 | 2003 | -1.11395 | 2.43166 | -5.87992 | 3.65203 | |
| 105 | 2004 | -0.14342 | 2.58740 | -5.21463 | 4.92779 | |
| y2 | 101 | 2000 | -2.09873 | 1.19096 | -4.43298 | 0.23551 |
| 102 | 2001 | -2.77050 | 1.47666 | -5.66469 | 0.12369 | |
| 103 | 2002 | -2.75724 | 1.74212 | -6.17173 | 0.65725 | |
| 104 | 2003 | -2.24943 | 2.01925 | -6.20709 | 1.70823 | |
| 105 | 2004 | -1.47460 | 2.25169 | -5.88782 | 2.93863 | |