Example 26.30 Linear Relations among Factor Loadings: COSAN Model Specification
This example reanalyzes the models in Example 26.26 by using the COSAN modeling language. The correlation matrix of six variables from Kinzer and Kinzer (N=326) is used (see Guttman 1957). McDonald (1980) uses this data set to demonstrate the fitting of a factor analysis model with linear constraints on factor loadings. Two factors are assumed for the data. The factor loading matrix 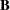 is shown in the following:
is shown in the following:
The loadings on the second factor are linearly related to the loadings on the first factor, as described by the following formula:
The correlation structures are represented by
where 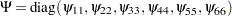 represents the diagonal matrix of unique variances for the variables. Because matrix
represents the diagonal matrix of unique variances for the variables. Because matrix 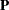 is a correlation matrix, its diagonal elements are fixed constants 1. This means that the diagonal elements of the correlation structures must also satisfy the following condition:
is a correlation matrix, its diagonal elements are fixed constants 1. This means that the diagonal elements of the correlation structures must also satisfy the following condition:
To analyze the correlation structures by using PROC CALIS, you formulate a covariance structure model with such correlation structures embedded in the model. That is, you want to fit the following covariance structure model to the Kinzer data:
where 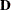 is a 6 x 6 diagonal matrix that contains the population standard deviations of the observed variables.
is a 6 x 6 diagonal matrix that contains the population standard deviations of the observed variables.
The following statements use the COSAN modeling language to specify this covariance structure model:
proc calis data=Kinzer nobs=326 nose;
cosan
var= var1-var6,
D(6,DIA) * B(2,GEN) + D(6,DIA) * Psi(6,DIA);
matrix B
[ ,1] = b11 b21 b31 b41 b51 b61,
[ ,2] = b12 b22 b32 b42 b52 b62;
matrix Psi
[1,1] = psi1-psi6;
matrix D
[1,1] = d1-d6;
parameters alpha (1.);
/* SAS Programming Statements to Define Dependent Parameters*/
/* 6 constraints on the factor loadings */
b12 = alpha - b11;
b22 = alpha - b21;
b32 = alpha - b31;
b42 = alpha - b41;
b52 = alpha - b51;
b62 = alpha - b61;
/* 6 Constraints on Correlation structures */
psi1 = 1. - b11 * b11 - b12 * b12;
psi2 = 1. - b21 * b21 - b22 * b22;
psi3 = 1. - b31 * b31 - b32 * b32;
psi4 = 1. - b41 * b41 - b42 * b42;
psi5 = 1. - b51 * b51 - b52 * b52;
psi6 = 1. - b61 * b61 - b62 * b62;
vnames
D = [var1-var6],
B = [factor1 factor2],
Psi = D;
run;
In the PROC CALIS statement, you specify the data set by the DATA= option and the number of observations by the NOBS= option. You also use the NOSE option to suppress the printing of the standard error estimates.
In the COSAN statement, you specify the variables for the covariance structure analysis in the VAR= option. Next, you specify the covariance structure formula for the variables. When generating the covariance structure expressions for the terms, PROC CALIS examines the matrix type of the last matrix in each term to determine how the expression is generated. If the last matrix in a term is not a symmetric matrix (including diagonal or identity matrix), the transpose of the last matrix would be included in the expression. This ensures that a symmetric matrix expression is formed for the covariance structures. For example, the first term in the current covariance structure formula is D(6,DIA)*B(2,GEN). Because 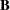 is not a symmetric matrix, the expression generated by PROC CALIS is
is not a symmetric matrix, the expression generated by PROC CALIS is
However, for the second term D(6,DIA)* Psi(6,DIA), matrix 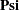 is a symmetric matrix so that the expression generated by PROC CALIS is
is a symmetric matrix so that the expression generated by PROC CALIS is
Output 26.30.1 shows the covariance structure model and the model matrices. With 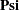 representing the unique variance matrix
representing the unique variance matrix 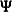 , the printed covariance structure formula for
, the printed covariance structure formula for 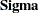 is clearly what you intend to specify.
is clearly what you intend to specify.
Output 26.30.1
The Covariance Structures and Model Matrices: Linearly Constrained Loadings
| Sigma = |
D*B*B`*D` + D*Psi*D` |
| B |
6 |
2 |
GEN: Rectangular |
| D |
6 |
6 |
DIA: Diagonal |
| Psi |
6 |
6 |
DIA: Diagonal |
In the MATRIX statements, you specify the parameters in the model matrices. You use parameters with the b prefix to name the two columns of loadings of the factor matrix 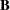 . You use free parameters psi1–psi6 for the diagonal elements of the
. You use free parameters psi1–psi6 for the diagonal elements of the 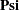 matrix, and free parameters d1–d6 for the diagonal elements of the
matrix, and free parameters d1–d6 for the diagonal elements of the 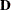 matrix. Next, you use a PARAMETERS statement to define an independent parameter alpha in the model. This parameter takes an initial value of 1.0. Using this independent parameter and six SAS programming statements, you define the loadings in the second column of matrix
matrix. Next, you use a PARAMETERS statement to define an independent parameter alpha in the model. This parameter takes an initial value of 1.0. Using this independent parameter and six SAS programming statements, you define the loadings in the second column of matrix 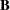 as functions of the loadings in the first column of the same matrix.
as functions of the loadings in the first column of the same matrix.
You use six more SAS programming statements to define the unique variance parameters psi1–psi6 as dependent parameters of the factor loadings. These constraints ensure that the embedded correlation structures have diagonal elements fixed at 1.0.
Lastly, you use the VNAMES statement to label the column names of the model matrices. The column names of the diagonal matrix 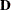 are the same as the observed variables. The column names of matrix
are the same as the observed variables. The column names of matrix 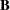 are for the factor names.
are for the factor names.
As compared with the covariance structure specification (that is, the second specification) by the LINEQS model in Example 26.26, the current COSAN specification seems to be more direct and concise in specifying the parameter constraints. Because of the direct references to the matrix elements in the COSAN modeling language, you can set the required 12 constraints in a very straightforward way as the 12 SAS programming statements in the preceding specification. However, with the LINEQS model specification language in Example 26.26, you need 18 more SAS programming statements to define the correct constraints for the same covariance structure model.
Output 26.30.2 shows the fit summary table. The chi-square test statistic is 14.63 with 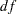 =8 (
=8 (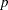 = 0.067). These are the same model fitting results as using the LINEQS model specification, as shown in Output 26.26.4 of Example 26.26.
= 0.067). These are the same model fitting results as using the LINEQS model specification, as shown in Output 26.26.4 of Example 26.26.
Output 26.30.2
Model Fit: Linearly Constrained Loadings with Embedded Correlation Structures
Output 26.30.3 shows the estimation of the loading matrix 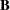 . These estimates of factor loadings are essentially the same as those obtained from the LINEQS model specification, as shown in Output 26.26.6, except that the two columns of the loading matrix
. These estimates of factor loadings are essentially the same as those obtained from the LINEQS model specification, as shown in Output 26.26.6, except that the two columns of the loading matrix 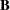 are switched. The column switching is not a concern because the factor labels are arbitrary.
are switched. The column switching is not a concern because the factor labels are arbitrary.
Output 26.30.3
Estimation of the 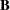 Matrix by the COSAN Model Specification
Matrix by the COSAN Model Specification
Output 26.30.4 shows the estimation of the scaling matrix 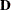 . All these standard deviation estimates for the observed variables match those obtained from the LINEQS model specification, as shown in Output 26.26.6.
. All these standard deviation estimates for the observed variables match those obtained from the LINEQS model specification, as shown in Output 26.26.6.
Output 26.30.4
Estimation of the 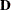 Matrix by the COSAN Model Specification
Matrix by the COSAN Model Specification
Output 26.30.5 shows the estimation of the unique covariance matrix 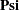 . All these unique variance parameter estimates match those obtained from the LINEQS model specification, as shown in Output 26.26.6.
. All these unique variance parameter estimates match those obtained from the LINEQS model specification, as shown in Output 26.26.6.
Output 26.30.5
Estimation of the 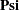 Matrix by the COSAN Model Specification
Matrix by the COSAN Model Specification
Finally, Output 26.30.6 shows the estimation of the independent parameter alpha. The same estimate of alpha is shown in Output 26.26.6.
Output 26.30.6
Estimation of the Independent Parameter alpha by the COSAN Model Specification


 Classes of Statements in PROC CALIS Single-Group Analysis Syntax Multiple-Group Multiple-Model Analysis Syntax PROC CALIS Statement BOUNDS Statement BY Statement COSAN Statement COV Statement DETERM Statement EFFPART Statement FACTOR Statement FITINDEX Statement FREQ Statement GROUP Statement LINCON Statement LINEQS Statement LISMOD Statement LMTESTS Statement MATRIX Statement MEAN Statement MODEL Statement MSTRUCT Statement NLINCON Statement NLOPTIONS Statement OUTFILES Statement PARAMETERS Statement PARTIAL Statement PATH Statement PCOV Statement PVAR Statement RAM Statement REFMODEL Statement RENAMEPARM Statement SAS Programming Statements SIMTESTS Statement STD Statement STRUCTEQ Statement TESTFUNC Statement VAR Statement VARIANCE Statement VARNAMES Statement WEIGHT Statement
Classes of Statements in PROC CALIS Single-Group Analysis Syntax Multiple-Group Multiple-Model Analysis Syntax PROC CALIS Statement BOUNDS Statement BY Statement COSAN Statement COV Statement DETERM Statement EFFPART Statement FACTOR Statement FITINDEX Statement FREQ Statement GROUP Statement LINCON Statement LINEQS Statement LISMOD Statement LMTESTS Statement MATRIX Statement MEAN Statement MODEL Statement MSTRUCT Statement NLINCON Statement NLOPTIONS Statement OUTFILES Statement PARAMETERS Statement PARTIAL Statement PATH Statement PCOV Statement PVAR Statement RAM Statement REFMODEL Statement RENAMEPARM Statement SAS Programming Statements SIMTESTS Statement STD Statement STRUCTEQ Statement TESTFUNC Statement VAR Statement VARIANCE Statement VARNAMES Statement WEIGHT Statement Input Data Sets Output Data Sets The COSAN Model The FACTOR Model The LINEQS Model The LISMOD Model and Submodels The MSTRUCT Model The PATH Model The RAM Model Naming Variables and Parameters Setting Constraints on Parameters Automatic Variable Selection Estimation Criteria Relationships among Estimation Criteria Gradient, Hessian, Information Matrix, and Approximate Standard Errors Counting the Degrees of Freedom Assessment of Fit Total, Direct, and Indirect Effects Standardized Solutions Modification Indices Missing Values and the Analysis of Missing Patterns Measures of Multivariate Kurtosis Initial Estimates Use of Optimization Techniques Computational Problems Displayed Output ODS Table Names ODS Graphics
Input Data Sets Output Data Sets The COSAN Model The FACTOR Model The LINEQS Model The LISMOD Model and Submodels The MSTRUCT Model The PATH Model The RAM Model Naming Variables and Parameters Setting Constraints on Parameters Automatic Variable Selection Estimation Criteria Relationships among Estimation Criteria Gradient, Hessian, Information Matrix, and Approximate Standard Errors Counting the Degrees of Freedom Assessment of Fit Total, Direct, and Indirect Effects Standardized Solutions Modification Indices Missing Values and the Analysis of Missing Patterns Measures of Multivariate Kurtosis Initial Estimates Use of Optimization Techniques Computational Problems Displayed Output ODS Table Names ODS Graphics Estimating Covariances and Correlations Estimating Covariances and Means Simultaneously Testing Uncorrelatedness of Variables Testing Covariance Patterns Testing Some Standard Covariance Pattern Hypotheses Linear Regression Model Multivariate Regression Models Measurement Error Models Testing Specific Measurement Error Models Measurement Error Models with Multiple Predictors Measurement Error Models Specified As Linear Equations Confirmatory Factor Models Confirmatory Factor Models: Some Variations The Full Information Maximum Likelihood Method Comparing the ML and FIML Estimation Path Analysis: Stability of Alienation Simultaneous Equations with Mean Structures and Reciprocal Paths Fitting Direct Covariance Structures Confirmatory Factor Analysis: Cognitive Abilities Testing Equality of Two Covariance Matrices Using a Multiple-Group Analysis Testing Equality of Covariance and Mean Matrices between Independent Groups Illustrating Various General Modeling Languages Testing Competing Path Models for the Career Aspiration Data Fitting a Latent Growth Curve Model Higher-Order and Hierarchical Factor Models Linear Relations among Factor Loadings Multiple-Group Model for Purchasing Behavior Fitting the RAM and EQS Models by the COSAN Modeling Language Second-Order Confirmatory Factor Analysis Linear Relations among Factor Loadings: COSAN Model Specification Ordinal Relations among Factor Loadings Longitudinal Factor Analysis
Estimating Covariances and Correlations Estimating Covariances and Means Simultaneously Testing Uncorrelatedness of Variables Testing Covariance Patterns Testing Some Standard Covariance Pattern Hypotheses Linear Regression Model Multivariate Regression Models Measurement Error Models Testing Specific Measurement Error Models Measurement Error Models with Multiple Predictors Measurement Error Models Specified As Linear Equations Confirmatory Factor Models Confirmatory Factor Models: Some Variations The Full Information Maximum Likelihood Method Comparing the ML and FIML Estimation Path Analysis: Stability of Alienation Simultaneous Equations with Mean Structures and Reciprocal Paths Fitting Direct Covariance Structures Confirmatory Factor Analysis: Cognitive Abilities Testing Equality of Two Covariance Matrices Using a Multiple-Group Analysis Testing Equality of Covariance and Mean Matrices between Independent Groups Illustrating Various General Modeling Languages Testing Competing Path Models for the Career Aspiration Data Fitting a Latent Growth Curve Model Higher-Order and Hierarchical Factor Models Linear Relations among Factor Loadings Multiple-Group Model for Purchasing Behavior Fitting the RAM and EQS Models by the COSAN Modeling Language Second-Order Confirmatory Factor Analysis Linear Relations among Factor Loadings: COSAN Model Specification Ordinal Relations among Factor Loadings Longitudinal Factor Analysis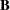 is shown in the following:
is shown in the following: 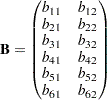
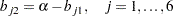
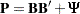
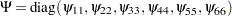 represents the diagonal matrix of unique variances for the variables. Because matrix
represents the diagonal matrix of unique variances for the variables. Because matrix 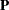 is a correlation matrix, its diagonal elements are fixed constants 1. This means that the diagonal elements of the correlation structures must also satisfy the following condition:
is a correlation matrix, its diagonal elements are fixed constants 1. This means that the diagonal elements of the correlation structures must also satisfy the following condition: 

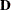 is a 6 x 6 diagonal matrix that contains the population standard deviations of the observed variables.
is a 6 x 6 diagonal matrix that contains the population standard deviations of the observed variables. 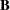 is not a symmetric matrix, the expression generated by PROC CALIS is
is not a symmetric matrix, the expression generated by PROC CALIS is 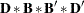
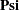 is a symmetric matrix so that the expression generated by PROC CALIS is
is a symmetric matrix so that the expression generated by PROC CALIS is 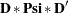
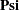 representing the unique variance matrix
representing the unique variance matrix 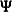 , the printed covariance structure formula for
, the printed covariance structure formula for 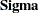 is clearly what you intend to specify.
is clearly what you intend to specify. 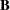 . You use free parameters psi1–psi6 for the diagonal elements of the
. You use free parameters psi1–psi6 for the diagonal elements of the 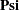 matrix, and free parameters d1–d6 for the diagonal elements of the
matrix, and free parameters d1–d6 for the diagonal elements of the 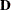 matrix. Next, you use a PARAMETERS statement to define an independent parameter alpha in the model. This parameter takes an initial value of 1.0. Using this independent parameter and six SAS programming statements, you define the loadings in the second column of matrix
matrix. Next, you use a PARAMETERS statement to define an independent parameter alpha in the model. This parameter takes an initial value of 1.0. Using this independent parameter and six SAS programming statements, you define the loadings in the second column of matrix 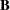 as functions of the loadings in the first column of the same matrix.
as functions of the loadings in the first column of the same matrix. 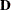 are the same as the observed variables. The column names of matrix
are the same as the observed variables. The column names of matrix 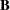 are for the factor names.
are for the factor names. 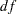 =8 (
=8 (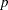 = 0.067). These are the same model fitting results as using the LINEQS model specification, as shown in Output 26.26.4 of Example 26.26.
= 0.067). These are the same model fitting results as using the LINEQS model specification, as shown in Output 26.26.4 of Example 26.26. 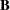 . These estimates of factor loadings are essentially the same as those obtained from the LINEQS model specification, as shown in Output 26.26.6, except that the two columns of the loading matrix
. These estimates of factor loadings are essentially the same as those obtained from the LINEQS model specification, as shown in Output 26.26.6, except that the two columns of the loading matrix 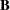 are switched. The column switching is not a concern because the factor labels are arbitrary.
are switched. The column switching is not a concern because the factor labels are arbitrary. 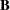 Matrix by the COSAN Model Specification
Matrix by the COSAN Model Specification
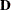 . All these standard deviation estimates for the observed variables match those obtained from the LINEQS model specification, as shown in Output 26.26.6.
. All these standard deviation estimates for the observed variables match those obtained from the LINEQS model specification, as shown in Output 26.26.6. 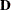 Matrix by the COSAN Model Specification
Matrix by the COSAN Model Specification
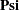 . All these unique variance parameter estimates match those obtained from the LINEQS model specification, as shown in Output 26.26.6.
. All these unique variance parameter estimates match those obtained from the LINEQS model specification, as shown in Output 26.26.6.  Matrix by the COSAN Model Specification
Matrix by the COSAN Model Specification