The GLM Procedure
-
Overview

-
Getting Started

-
Syntax

-
Details
 Statistical Assumptions for Using PROC GLMSpecification of EffectsUsing PROC GLM InteractivelyParameterization of PROC GLM ModelsHypothesis Testing in PROC GLMEffect Size Measures for F Tests in GLMAbsorptionSpecification of ESTIMATE ExpressionsComparing GroupsMultivariate Analysis of VarianceRepeated Measures Analysis of VarianceRandom-Effects AnalysisMissing ValuesComputational ResourcesComputational MethodOutput Data SetsDisplayed OutputODS Table NamesODS Graphics
Statistical Assumptions for Using PROC GLMSpecification of EffectsUsing PROC GLM InteractivelyParameterization of PROC GLM ModelsHypothesis Testing in PROC GLMEffect Size Measures for F Tests in GLMAbsorptionSpecification of ESTIMATE ExpressionsComparing GroupsMultivariate Analysis of VarianceRepeated Measures Analysis of VarianceRandom-Effects AnalysisMissing ValuesComputational ResourcesComputational MethodOutput Data SetsDisplayed OutputODS Table NamesODS Graphics -
Examples
 Randomized Complete Blocks with Means Comparisons and ContrastsRegression with Mileage DataUnbalanced ANOVA for Two-Way Design with InteractionAnalysis of CovarianceThree-Way Analysis of Variance with ContrastsMultivariate Analysis of VarianceRepeated Measures Analysis of VarianceMixed Model Analysis of Variance with the RANDOM StatementAnalyzing a Doubly Multivariate Repeated Measures DesignTesting for Equal Group VariancesAnalysis of a Screening Design
Randomized Complete Blocks with Means Comparisons and ContrastsRegression with Mileage DataUnbalanced ANOVA for Two-Way Design with InteractionAnalysis of CovarianceThree-Way Analysis of Variance with ContrastsMultivariate Analysis of VarianceRepeated Measures Analysis of VarianceMixed Model Analysis of Variance with the RANDOM StatementAnalyzing a Doubly Multivariate Repeated Measures DesignTesting for Equal Group VariancesAnalysis of a Screening Design - References
If you fit several dependent variables to the same effects, you might want to make joint tests involving parameters of several dependent variables. Suppose you have p dependent variables, k parameters for each dependent variable, and n observations. The models can be collected into one equation:
where ![]() is
is ![]() ,
, ![]() is
is ![]() ,
, ![]() is
is ![]() , and
, and ![]() is
is ![]() . Each of the p models can be estimated and tested separately. However, you might also want to consider the joint distribution and test the
p models simultaneously.
. Each of the p models can be estimated and tested separately. However, you might also want to consider the joint distribution and test the
p models simultaneously.
For multivariate tests, you need to make some assumptions about the errors. With p dependent variables, there are ![]() errors that are independent across observations but not across dependent variables. Assume
errors that are independent across observations but not across dependent variables. Assume
where vec![]() strings
strings ![]() out by rows,
out by rows, ![]() denotes Kronecker product multiplication, and
denotes Kronecker product multiplication, and ![]() is
is ![]() .
. ![]() can be estimated by
can be estimated by
where ![]() , r is the rank of the
, r is the rank of the ![]() matrix, and
matrix, and ![]() is the matrix of residuals.
is the matrix of residuals.
If ![]() is scaled to unit diagonals, the values in
is scaled to unit diagonals, the values in ![]() are called partial correlations of the Ys adjusting for the Xs. This matrix can be displayed by PROC GLM if PRINTE
is specified as a MANOVA
option.
are called partial correlations of the Ys adjusting for the Xs. This matrix can be displayed by PROC GLM if PRINTE
is specified as a MANOVA
option.
The multivariate general linear hypothesis is written
You can form hypotheses for linear combinations across columns, as well as across rows of ![]() .
.
The MANOVA
statement of the GLM procedure tests special cases where ![]() corresponds to Type I, Type II, Type III, or Type IV tests, and
corresponds to Type I, Type II, Type III, or Type IV tests, and ![]() is the
is the ![]() identity matrix. These tests are joint tests that the given type of hypothesis holds for all dependent variables in the model,
and they are often sufficient to test all hypotheses of interest.
identity matrix. These tests are joint tests that the given type of hypothesis holds for all dependent variables in the model,
and they are often sufficient to test all hypotheses of interest.
Finally, when these special cases are not appropriate, you can specify your own ![]() and
and ![]() matrices by using the CONTRAST
statement before the MANOVA
statement and the M=
specification in the MANOVA
statement, respectively. Another alternative is to use a REPEATED
statement, which automatically generates a variety of
matrices by using the CONTRAST
statement before the MANOVA
statement and the M=
specification in the MANOVA
statement, respectively. Another alternative is to use a REPEATED
statement, which automatically generates a variety of ![]() matrices useful in repeated measures analysis of variance. See the section REPEATED Statement and the section Repeated Measures Analysis of Variance for more information.
matrices useful in repeated measures analysis of variance. See the section REPEATED Statement and the section Repeated Measures Analysis of Variance for more information.
One useful way to think of a MANOVA analysis with an ![]() matrix other than the identity is as an analysis of a set of transformed variables defined by the columns of the
matrix other than the identity is as an analysis of a set of transformed variables defined by the columns of the ![]() matrix. You should note, however, that PROC GLM always displays the
matrix. You should note, however, that PROC GLM always displays the ![]() matrix in such a way that the transformed variables are defined by the rows, not the columns, of the displayed
matrix in such a way that the transformed variables are defined by the rows, not the columns, of the displayed ![]() matrix.
matrix.
All multivariate tests carried out by the GLM procedure first construct the matrices ![]() and
and ![]() corresponding to the numerator and denominator, respectively, of a univariate F test:
corresponding to the numerator and denominator, respectively, of a univariate F test:
The diagonal elements of ![]() and
and ![]() correspond to the hypothesis and error SS for univariate tests. When the
correspond to the hypothesis and error SS for univariate tests. When the ![]() matrix is the identity matrix (the default), these tests are for the original dependent variables on the left side of the
MODEL
statement. When an
matrix is the identity matrix (the default), these tests are for the original dependent variables on the left side of the
MODEL
statement. When an ![]() matrix other than the identity is specified, the tests are for transformed variables defined by the columns of the
matrix other than the identity is specified, the tests are for transformed variables defined by the columns of the ![]() matrix. These tests can be studied by requesting the SUMMARY option, which produces univariate analyses for each original
or transformed variable.
matrix. These tests can be studied by requesting the SUMMARY option, which produces univariate analyses for each original
or transformed variable.
Four multivariate test statistics, all functions of the eigenvalues of ![]() (or
(or ![]() ), are constructed:
), are constructed:
-
Wilks’ lambda = det
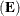 /det
/det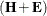
-
Pillai’s trace = trace
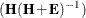
-
Hotelling-Lawley trace = trace
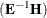
-
Roy’s greatest root =
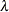 , largest eigenvalue of
, largest eigenvalue of 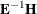
By default, all four are reported with p-values based on F approximations, as discussed in the "Multivariate Tests" section in Chapter 4: Introduction to Regression Procedures. Alternatively, if you specify MSTAT=EXACT in the associated MANOVA or REPEATED statement, p-values for three of the four tests are computed exactly (Wilks’ lambda, the Hotelling-Lawley trace, and Roy’s greatest root), and the p-values for the fourth (Pillai’s trace) are based on an F approximation that is more accurate than the default. See the "Multivariate Tests" section in Chapter 4: Introduction to Regression Procedures, for more details on the exact calculations.