The CALIS Procedure
-
Overview

-
Getting Started

-
Syntax
 Classes of Statements in PROC CALISSingle-Group Analysis SyntaxMultiple-Group Multiple-Model Analysis SyntaxPROC CALIS StatementBOUNDS StatementBY StatementCOSAN StatementCOV StatementDETERM StatementEFFPART StatementFACTOR StatementFITINDEX StatementFREQ StatementGROUP StatementLINCON StatementLINEQS StatementLISMOD StatementLMTESTS StatementMATRIX StatementMEAN StatementMODEL StatementMSTRUCT StatementNLINCON StatementNLOPTIONS StatementOUTFILES StatementPARAMETERS StatementPARTIAL StatementPATH StatementPATHDIAGRAM StatementPCOV StatementPVAR StatementRAM StatementREFMODEL StatementRENAMEPARM StatementSAS Programming StatementsSIMTESTS StatementSTD StatementSTRUCTEQ StatementTESTFUNC StatementVAR StatementVARIANCE StatementVARNAMES StatementWEIGHT Statement
Classes of Statements in PROC CALISSingle-Group Analysis SyntaxMultiple-Group Multiple-Model Analysis SyntaxPROC CALIS StatementBOUNDS StatementBY StatementCOSAN StatementCOV StatementDETERM StatementEFFPART StatementFACTOR StatementFITINDEX StatementFREQ StatementGROUP StatementLINCON StatementLINEQS StatementLISMOD StatementLMTESTS StatementMATRIX StatementMEAN StatementMODEL StatementMSTRUCT StatementNLINCON StatementNLOPTIONS StatementOUTFILES StatementPARAMETERS StatementPARTIAL StatementPATH StatementPATHDIAGRAM StatementPCOV StatementPVAR StatementRAM StatementREFMODEL StatementRENAMEPARM StatementSAS Programming StatementsSIMTESTS StatementSTD StatementSTRUCTEQ StatementTESTFUNC StatementVAR StatementVARIANCE StatementVARNAMES StatementWEIGHT Statement -
Details
 Input Data SetsOutput Data SetsDefault Analysis Type and Default ParameterizationThe COSAN ModelThe FACTOR ModelThe LINEQS ModelThe LISMOD Model and SubmodelsThe MSTRUCT ModelThe PATH ModelThe RAM ModelNaming Variables and ParametersSetting Constraints on ParametersAutomatic Variable SelectionPath Diagrams: Layout Algorithms, Default Settings, and CustomizationEstimation CriteriaRelationships among Estimation CriteriaGradient, Hessian, Information Matrix, and Approximate Standard ErrorsCounting the Degrees of FreedomAssessment of FitCase-Level Residuals, Outliers, Leverage Observations, and Residual DiagnosticsTotal, Direct, and Indirect EffectsStandardized SolutionsModification IndicesMissing Values and the Analysis of Missing PatternsMeasures of Multivariate KurtosisInitial EstimatesUse of Optimization TechniquesComputational ProblemsDisplayed OutputODS Table NamesODS Graphics
Input Data SetsOutput Data SetsDefault Analysis Type and Default ParameterizationThe COSAN ModelThe FACTOR ModelThe LINEQS ModelThe LISMOD Model and SubmodelsThe MSTRUCT ModelThe PATH ModelThe RAM ModelNaming Variables and ParametersSetting Constraints on ParametersAutomatic Variable SelectionPath Diagrams: Layout Algorithms, Default Settings, and CustomizationEstimation CriteriaRelationships among Estimation CriteriaGradient, Hessian, Information Matrix, and Approximate Standard ErrorsCounting the Degrees of FreedomAssessment of FitCase-Level Residuals, Outliers, Leverage Observations, and Residual DiagnosticsTotal, Direct, and Indirect EffectsStandardized SolutionsModification IndicesMissing Values and the Analysis of Missing PatternsMeasures of Multivariate KurtosisInitial EstimatesUse of Optimization TechniquesComputational ProblemsDisplayed OutputODS Table NamesODS Graphics -
Examples
 Estimating Covariances and CorrelationsEstimating Covariances and Means SimultaneouslyTesting Uncorrelatedness of VariablesTesting Covariance PatternsTesting Some Standard Covariance Pattern HypothesesLinear Regression ModelMultivariate Regression ModelsMeasurement Error ModelsTesting Specific Measurement Error ModelsMeasurement Error Models with Multiple PredictorsMeasurement Error Models Specified As Linear EquationsConfirmatory Factor ModelsConfirmatory Factor Models: Some VariationsResidual Diagnostics and Robust EstimationThe Full Information Maximum Likelihood MethodComparing the ML and FIML EstimationPath Analysis: Stability of AlienationSimultaneous Equations with Mean Structures and Reciprocal PathsFitting Direct Covariance StructuresConfirmatory Factor Analysis: Cognitive AbilitiesTesting Equality of Two Covariance Matrices Using a Multiple-Group AnalysisTesting Equality of Covariance and Mean Matrices between Independent GroupsIllustrating Various General Modeling LanguagesTesting Competing Path Models for the Career Aspiration DataFitting a Latent Growth Curve ModelHigher-Order and Hierarchical Factor ModelsLinear Relations among Factor LoadingsMultiple-Group Model for Purchasing BehaviorFitting the RAM and EQS Models by the COSAN Modeling LanguageSecond-Order Confirmatory Factor AnalysisLinear Relations among Factor Loadings: COSAN Model SpecificationOrdinal Relations among Factor LoadingsLongitudinal Factor Analysis
Estimating Covariances and CorrelationsEstimating Covariances and Means SimultaneouslyTesting Uncorrelatedness of VariablesTesting Covariance PatternsTesting Some Standard Covariance Pattern HypothesesLinear Regression ModelMultivariate Regression ModelsMeasurement Error ModelsTesting Specific Measurement Error ModelsMeasurement Error Models with Multiple PredictorsMeasurement Error Models Specified As Linear EquationsConfirmatory Factor ModelsConfirmatory Factor Models: Some VariationsResidual Diagnostics and Robust EstimationThe Full Information Maximum Likelihood MethodComparing the ML and FIML EstimationPath Analysis: Stability of AlienationSimultaneous Equations with Mean Structures and Reciprocal PathsFitting Direct Covariance StructuresConfirmatory Factor Analysis: Cognitive AbilitiesTesting Equality of Two Covariance Matrices Using a Multiple-Group AnalysisTesting Equality of Covariance and Mean Matrices between Independent GroupsIllustrating Various General Modeling LanguagesTesting Competing Path Models for the Career Aspiration DataFitting a Latent Growth Curve ModelHigher-Order and Hierarchical Factor ModelsLinear Relations among Factor LoadingsMultiple-Group Model for Purchasing BehaviorFitting the RAM and EQS Models by the COSAN Modeling LanguageSecond-Order Confirmatory Factor AnalysisLinear Relations among Factor Loadings: COSAN Model SpecificationOrdinal Relations among Factor LoadingsLongitudinal Factor Analysis - References
For a single-sample setting with a discrepancy function ![]() , the gradient is defined as the first partial derivatives of the discrepancy function with respect to the model parameters
, the gradient is defined as the first partial derivatives of the discrepancy function with respect to the model parameters
![]() :
:
The Hessian is defined as the second partial derivatives of the discrepancy function with respect to the model parameters
![]() :
:
Suppose that the mean and covariance structures fit perfectly with ![]() in the population. The information matrix is defined as
in the population. The information matrix is defined as
where the expectation ![]() is taken over the sampling space of
is taken over the sampling space of ![]() .
.
The information matrix plays a significant role in statistical theory. Under certain regularity conditions, the inverse of
the information matrix ![]() is the asymptotic covariance matrix for
is the asymptotic covariance matrix for ![]() , where N denotes the sample size and
, where N denotes the sample size and ![]() is an estimator.
is an estimator.
In practice, ![]() is never known and can only be estimated. The information matrix is therefore estimated by the so-called empirical information
matrix:
is never known and can only be estimated. The information matrix is therefore estimated by the so-called empirical information
matrix:
which is evaluated at the values of the sample estimates ![]() . Notice that this empirical information matrix, rather than the unknown
. Notice that this empirical information matrix, rather than the unknown ![]() , is the “information matrix” displayed in PROC CALIS output.
, is the “information matrix” displayed in PROC CALIS output.
Taking the inverse of the empirical information matrix with sample size adjustment, PROC CALIS approximates the estimated
covariance matrix of ![]() by:
by:
Approximate standard errors for ![]() can then be computed as the square roots of the diagonal elements of the estimated covariance matrix. The theory about the
empirical information matrix, the approximate covariance matrix of the parameter estimates, and the approximate standard errors
applies to all but the ULS and DWLS estimation methods. Standard errors are therefore not computed with the ULS and DWLS estimation
methods.
can then be computed as the square roots of the diagonal elements of the estimated covariance matrix. The theory about the
empirical information matrix, the approximate covariance matrix of the parameter estimates, and the approximate standard errors
applies to all but the ULS and DWLS estimation methods. Standard errors are therefore not computed with the ULS and DWLS estimation
methods.
If a given Hessian or information matrix is singular, PROC CALIS offers two ways to compute a generalized inverse of the matrix and, therefore, two ways to compute approximate standard errors of implicitly constrained parameter estimates, t values, and modification indices. Depending on the G4= specification, either a Moore-Penrose inverse or a G2 inverse is computed. The expensive Moore-Penrose inverse computes an estimate of the null space by using an eigenvalue decomposition. The cheaper G2 inverse is produced by sweeping the linearly independent rows and columns and zeroing out the dependent ones.
In the section Multiple-Group Discrepancy Function, the overall discrepancy function for multiple-group analysis is defined. The same notation is applied here. To begin with,
the overall discrepancy function ![]() is expressed as a weighted sum of individual discrepancy functions
is expressed as a weighted sum of individual discrepancy functions ![]() ’s for the groups as follows:
’s for the groups as follows:
where
is the weight for group i,
is the total sample size, and ![]() is the sample size for group i.
is the sample size for group i.
The gradient ![]() and the Hessian
and the Hessian ![]() are now defined as weighted sum of individual functions. That is,
are now defined as weighted sum of individual functions. That is,
and
Suppose that the mean and covariance structures fit perfectly with ![]() in the population. If each
in the population. If each ![]() converges to a fixed constant
converges to a fixed constant ![]() (
(![]() ) with increasing total sample size, the information matrix can be written as:
) with increasing total sample size, the information matrix can be written as:
To approximate this information matrix, an empirical counterpart is used:
which is evaluated at the values of the sample estimates ![]() . Again, this empirical information matrix, rather than the unknown
. Again, this empirical information matrix, rather than the unknown ![]() , is the “information matrix” output in PROC CALIS results.
, is the “information matrix” output in PROC CALIS results.
Taking the inverse of the empirical information matrix with sample size adjustment, PROC CALIS approximates the estimated
covariance matrix of ![]() in multiple-group analysis by:
in multiple-group analysis by:
Approximate standard errors for ![]() can then be computed as the square roots of the diagonal elements of the estimated covariance matrix. Again, for ULS and
DWLS estimation, the theory does not apply and so there are no standard errors computed in these cases.
can then be computed as the square roots of the diagonal elements of the estimated covariance matrix. Again, for ULS and
DWLS estimation, the theory does not apply and so there are no standard errors computed in these cases.
When computing the approximate covariance matrix and hence the standard errors for the parameter estimates, inversion of the scaled information matrix or Hessian matrix is involved. The numerical condition of the information matrix can be very poor in many practical applications, especially for the analysis of unscaled covariance data. The following four-step strategy is used for the inversion of the information matrix.
-
The inversion (usually of a normalized matrix
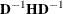 ) is tried using a modified form of the Bunch and Kaufman (1977) algorithm, which allows the specification of a different singularity criterion for each pivot. The following three criteria
for the detection of rank loss in the information matrix are used to specify thresholds:
) is tried using a modified form of the Bunch and Kaufman (1977) algorithm, which allows the specification of a different singularity criterion for each pivot. The following three criteria
for the detection of rank loss in the information matrix are used to specify thresholds:
-
ASING specifies absolute singularity.
-
MSING specifies relative singularity depending on the whole matrix norm.
-
VSING specifies relative singularity depending on the column matrix norm.
If no rank loss is detected, the inverse of the information matrix is used for the covariance matrix of parameter estimates, and the next two steps are skipped.
-
-
The linear dependencies among the parameter subsets are displayed based on the singularity criteria.
-
If the number of parameters t is smaller than the value specified by the G4= option (the default value is 60), the Moore-Penrose inverse is computed based on the eigenvalue decomposition of the information matrix. If you do not specify the NOPRINT option, the distribution of eigenvalues is displayed, and those eigenvalues that are set to zero in the Moore-Penrose inverse are indicated. You should inspect this eigenvalue distribution carefully.
-
If PROC CALIS did not set the right subset of eigenvalues to zero, you can specify the COVSING= option to set a larger or smaller subset of eigenvalues to zero in a further run of PROC CALIS.