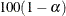-
AB < (ADJUST)>
-
requests an analysis of Ansari-Bradley scores.
For more information, see the section Ansari-Bradley Scores. The ADJUST option subtracts class medians from the input observations before performing the analysis.
-
ADJUST
-
adjusts for location differences among classes
before performing tests for scale differences. The ADJUST option applies to the following analyses: Ansari-Bradley (AB
), Klotz (KLOTZ
), Mood (MOOD
), and Siegel-Tukey (ST
). If you specify the ADJUST option, PROC NPAR1WAY subtracts the corresponding class median from each observation before scoring
the data and performing the scale tests that you request.
You can also request adjustment for an individual scale test by specifying ADJUST in parentheses after the scale test option;
for example, you can specify ST(ADJUST)
to adjust by class medians before performing the Siegel-Tukey test.
-
ALIGN=STRATA < (option)>
-
aligns the response variable values by strata (Hodges and Lehmann 1962; Mehrotra, Lu, and Li 2010) before performing the analyses that you request. When you specify this option, you must also specify a STRATA
statement to define the strata. PROC NPAR1WAY subtracts the stratum location measure from each response value in the stratum.
You can specify a location measure option in parentheses after the ALIGN=STRATA option; by default, PROC NPAR1WAY uses the stratum median (MEDIAN
) as the stratum location measure.
You can specify one of the following options:
-
HL
-
specifies the Hodges-Lehmann shift as the stratum location measure. The stratum Hodges-Lehmann shift is the median of all
pairwise (Walsh) averages of response values in the stratum.
-
MEAN
-
specifies the mean (average) of the response values in the stratum as the stratum location measure.
-
MEDIAN
-
specifies the median of the response values in the stratum as the stratum location measure. This is the default location measure
for ALIGN=STRATA.
-
ALPHA=

-
specifies the level of the confidence limits for the Hodges-Lehmann location shift, which
you can request by specifying the HL
option. The value of  must be between 0 and 1; a confidence level of
must be between 0 and 1; a confidence level of  produces
produces 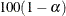 % confidence limits. By default, ALPHA=0.05, which produces 95% confidence limits for the location shift.
% confidence limits. By default, ALPHA=0.05, which produces 95% confidence limits for the location shift.
-
ANOVA
-
requests a standard analysis of variance on the raw data.
-
CONOVER
-
requests an analysis of Conover scores.
For more information, see the section Conover Scores.
-
CORRECT=NO
-
suppresses the continuity correction for the Wilcoxon two-sample test
and the Siegel-Tukey two-sample test. For more information, see the section Continuity Correction.
-
D
-
requests the one-sided Kolmogorov-Smirnov  and
and  statistics
and their asymptotic p-values, in addition to the two-sided D statistic that the EDF
option produces for two-sample data. The D option invokes the EDF
option. If you request exact Kolmogorov-Smirnov tests by specifying the KS option in the EXACT
statement for two-sample data, PROC NPAR1WAY provides D+ and D– by default. For more information about Kolmogorov-Smirnov statistics, see the section Tests Based on the Empirical Distribution Function.
statistics
and their asymptotic p-values, in addition to the two-sided D statistic that the EDF
option produces for two-sample data. The D option invokes the EDF
option. If you request exact Kolmogorov-Smirnov tests by specifying the KS option in the EXACT
statement for two-sample data, PROC NPAR1WAY provides D+ and D– by default. For more information about Kolmogorov-Smirnov statistics, see the section Tests Based on the Empirical Distribution Function.
-
DATA=SAS-data-set
-
names the SAS-data-set to be analyzed by PROC NPAR1WAY. If you omit the
DATA= option, the procedure uses the most recently created SAS data set.
-
DSCF
-
requests the Dwass, Steel, Critchlow-Fligner multiple comparison
procedure, which is based on pairwise two-sample rankings. This procedure is available for multisample data, where the number
of CLASS
variable levels is greater than 2. For more information, see the section Multiple Comparisons Based on Pairwise Rankings.
-
EDF
KS
-
requests statistics based on the empirical distribution function. These
include the Kolmogorov-Smirnov and Cramér–von Mises tests and, if there are only two classification levels, the Kuiper test.
For more information, see the section Tests Based on the Empirical Distribution Function.
The EDF option produces the Kolmogorov-Smirnov D statistic for two-sample data. You can request the one-sided D+ and D– statistics for two-sample data by specifying the D
option.
-
FP < (REFCLASS=class-number | 'class-value')>
-
requests the Fligner-Policello test for two-sample data.
For more information, see the section Fligner-Policello Test.
The REFCLASS= option specifies which of the two CLASS
variable levels (samples) to use as the reference class X for the difference between class placement sums,  . REFCLASS=1 refers to the first class that is listed in the "Fligner-Policello Placements" table, and REFCLASS=2 refers to
the second class listed. The table displays class levels in the order in which they appear in the input data set. REFCLASS=class-value identifies the reference class by the formatted value of the CLASS
variable. By default, the reference class is the second class listed in the "Fligner-Policello Placements" table (REFCLASS=2).
. REFCLASS=1 refers to the first class that is listed in the "Fligner-Policello Placements" table, and REFCLASS=2 refers to
the second class listed. The table displays class levels in the order in which they appear in the input data set. REFCLASS=class-value identifies the reference class by the formatted value of the CLASS
variable. By default, the reference class is the second class listed in the "Fligner-Policello Placements" table (REFCLASS=2).
-
HL < (REFCLASS=class-number | 'class-value')>
-
requests Hodges-Lehmann estimation of the location shift for two-sample
data. This option also provides asymptotic confidence limits for the location shift (which are sometimes known as Moses confidence
limits). For more information, see the section Hodges-Lehmann Estimation of Location Shift. You can specify the confidence level in the ALPHA=
option. By default, ALPHA=0.05, which produces 95% confidence limits for the location shift.
The REFCLASS= option specifies which of the two CLASS
variable levels (samples) to use as the reference class X for the location shift, Y – X. REFCLASS=1 refers to the first class that is listed in the "Wilcoxon Scores" table, and REFCLASS=2 refers to the second
class listed. The table displays class levels in the order in which they appear in the input data set. REFCLASS=class-value identifies the reference class by the formatted value of the CLASS
variable.
By default, PROC NPAR1WAY uses the larger of the two classes as the reference class X to compute the Hodges-Lehmann location shift, Y – X. If both class have the same number of observations, PROC NPAR1WAY uses the class that appears second in the input data set
as the reference class.
-
KLOTZ < (ADJUST)>
-
requests an analysis of Klotz scores.
For more information, see the section Klotz Scores. If you specify the ADJUST option, PROC NPAR1WAY subtracts the corresponding class median from each observation before performing
the analysis.
-
MEDIAN
-
requests an analysis of median scores. When there are two classification
levels, this option produces the two-sample median test. When there are more than two samples, this option produces the multisample
median test, which is also known as the Brown-Mood test. For more information, see the section Median Scores.
-
MISSING
-
treats missing values as a valid nonmissing level
for CLASS
and STRATA
variables. By default, PROC NPAR1WAY excludes an observations from the analysis if the value of the CLASS variable is missing.
By default, PROC NPAR1WAY excludes an observations from the stratified analysis if the value of the STRATA variable is missing.
For more information, see the section Missing Values.
-
MOOD < (ADJUST)>
-
requests an analysis of Mood scores.
For more information, see the section Mood Scores. If you specify the ADJUST option, PROC NPAR1WAY subtracts the corresponding class median from each observation before performing
the analysis.
-
NOPRINT
-
suppresses the display of all output. You can use the NOPRINT option
when you only want to create an output data set. This option temporarily disables the Output Delivery System (ODS). For more
information, see Chapter 20: Using the Output Delivery System.
-
PLOTS < (global-plot-options)> < =plot-request < (plot-option)> >
PLOTS < (global-plot-options)> < =(plot-request < (plot-option)> < …plot-request < (plot-option)> > )>
-
controls the plots that are produced through ODS Graphics.
Plot-requests specify the plots to produce, and plot-options control the appearance and content of the plots. You can specify plot-options in parentheses after a plot-request. A global-plot-option applies to all plots for which it is available, unless it is altered by a specific plot-option. You can specify global-plot-options in parentheses after the PLOTS option.
When you specify only one plot-request, you can omit the parentheses around the request. For example:
plots=all
plots=wilcoxonboxplot
plots=(wilcoxonboxplot edfplot)
plots(only)=(medianplot normalboxplot)
ODS Graphics must be enabled before plots can be requested. For example:
ods graphics on;
proc npar1way plots=wilcoxonboxplot;
variable response;
class treatment;
run;
ods graphics off;
For more information about enabling and disabling ODS Graphics, see the section Enabling and Disabling ODS Graphics in Chapter 21: Statistical Graphics Using ODS.
If ODS Graphics is enabled but you do not specify the PLOTS= option, PROC NPAR1WAY produces all plots that are associated
with the analyses that you request. If you request a plot by specifying the PLOTS= option but do not request the corresponding
analysis, PROC NPAR1WAY automatically invokes that analysis. For example, if you specify PLOTS=CONOVERBOXPLOT but do not also
specify the CONOVER option in the PROC NPAR1WAY statement, PROC NPAR1WAY produces the Conover scores analysis in addition
to the box plot.
You can suppress default plots and request specific plots by using the PLOTS(ONLY)=
option; PLOTS(ONLY)=(plot-requests) produces only the plots that are specified as plot-requests. You can suppress all plots by specifying the PLOTS=NONE
option. The PLOTS= option has no effect when you specify the NOPRINT
option.
PROC NPAR1WAY provides box plots of scored data, median plots, and empirical distribution plots. See Figure 83.7, Output 83.1.2, Output 83.1.4, and Output 83.2.2 for examples of plots that PROC NPAR1WAY produces. For general information about ODS Graphics, see Chapter 21: Statistical Graphics Using ODS.
Global Plot Options
A global-plot-option applies to all plots for which the option is available unless it is altered by a specific plot-option. You can specify the following global-plot-options in parentheses after the PLOTS option. You cannot specify both the STATS and the NOSTATS global-plot-options in the same statement.
-
NOSTATS
-
suppresses the p-values that are displayed by default in the plots.
-
ONLY
-
suppresses the default plots and requests only the plots that are specified as plot-requests.
-
STATS
-
displays p-values in the plots. This is the default.
Plot Requests
You can specify the following plot-requests:
-
ABBOXPLOT
AB
-
requests a box plot of Ansari-Bradley scores. This plot is associated
with the Ansari-Bradley analysis, which you request by specifying the AB
option.
-
ALL
-
requests all plots that are associated with the specified analyses. This is the
default if you do not specify the ONLY
global-plot-option.
-
ANOVABOXPLOT
ANOVA
-
requests a box plot of the raw data. This plot is associated with the
analysis of variance based on the raw data, which you request by specifying the ANOVA
option.
-
CONOVERBOXPLOT
CONOVER
-
requests a box plot of Conover scores. This plot is associated
with the Conover analysis, which you request by specifying the CONOVER
option.
-
DATASCORESBOXPLOT
DATASCORES
-
requests a box plot of raw data scores. This plot is associated
with the analysis that uses input data as scores, which you request by specifying the SCORES=DATA
option.
-
EDFPLOT
EDF
-
requests an empirical distribution plot. This plot is associated with
the analyses based on the empirical distribution function, which you request by specifying the EDF
option.
-
FPBOXPLOT
FP
-
requests a box plot of Fligner-Policello placements. This plot is
associated with the Fligner-Policello analysis, which you request by specifying the FP
option.
-
KLOTZBOXPLOT
KLOTZ
-
requests a box plot of Klotz scores. This plot is associated with the
Klotz analysis, which you request by specifying the KLOTZ
option.
-
MEDIANPLOT
MEDIAN
-
requests a stacked bar chart showing the frequencies above and below
the overall median. This plot is associated with the median score analysis, which you request by specifying the MEDIAN
option.
-
MOODBOXPLOT
MOOD
-
requests a box plot of Mood scores. This plot is associated with the
Mood analysis, which you request by specifying the MOOD
option.
-
NONE
-
suppresses all plots.
-
SAVAGEBOXPLOT
SAVAGE
-
requests a box plot of Savage scores. This plot is associated with the
Savage analysis, which you request by specifying the SAVAGE
option.
-
STBOXPLOT
ST
-
requests a box plot of Siegel-Tukey scores. This plot is associated
with the Siegel-Tukey analysis, which you request by specifying the ST
option.
-
VWBOXPLOT
VW
NORMALBOXPLOT
NORMAL
-
requests a box plot of Van der Waerden (normal) scores. This plot is
associated with the Van der Waerden analysis, which you request by specifying the VW
(NORMAL) option.
-
WILCOXONBOXPLOT
WILCOXON
-
requests a box plot of Wilcoxon scores. This plot is associated with
the Wilcoxon analysis, which you request by specifying the WILCOXON
option.
Plot Options
The following plot-options are available for any plot-request. You cannot specify both the STATS and the NOSTATS plot-options for the same plot. If you specify both the NOSTATS global-plot-option and the STATS plot-option for an individual plot-request, the STATS plot-option overrides the global-plot-option and displays statistics in the individual plot.
-
NOSTATS
-
suppresses the p-values that are displayed in the plot by default.
-
STATS
-
displays p-values in the plot. This is the default.
-
SAVAGE
-
requests an analysis of Savage scores.
For more information, see the section Savage Scores.
-
SCORES=DATA
PERM
-
requests an analysis that uses input data as scores. This option gives
you the flexibility to construct any scores for your data with the DATA step and then analyze these scores with PROC NPAR1WAY.
For more information, see the section Scores for Linear Rank and One-Way ANOVA Tests.
To produce the two-sample permutation test that is known as Pitman’s test, provide raw (unscored) data in the input data set
and specify the SCORES=DATA option in the EXACT
statement. For more information, see the section Exact Tests.
-
ST < (ADJUST)>
-
requests an analysis of Siegel-Tukey scores.
For more information, see the section Siegel-Tukey Scores. If you specify the ADJUST option, PROC NPAR1WAY subtracts the corresponding class median from each observation before performing
the analysis.
-
VW
NORMAL
-
requests an analysis of Van der Waerden (normal) scores.
For more information, see the section Van der Waerden (Normal) Scores.
-
WILCOXON
-
requests an analysis of Wilcoxon scores. When there are two
classification levels (samples), this option produces the Wilcoxon rank-sum test. For any number of classification levels,
this option produces the Kruskal-Wallis test. For more information, see the section Wilcoxon Scores.