-
ADJRSQ <(adjrsq-options)>
-
displays the adjusted R-square values for the models examined when you request variable selection with the SELECTION= option
in the MODEL
statement.
The following adjrsq-options are available for models where you request the RSQUARE, ADJRSQ, or CP selection method:
-
LABEL
-
requests that the model number corresponding to the one displayed in the "Subset Selection Summary" table be used to label
the model with the largest adjusted R-square statistic at each value of the number of parameters.
-
LABELVARS
-
requests that the list (excluding the intercept) of the regressors in the relevant model be used to label the model with the
largest adjusted R-square statistic at each value of the number of parameters.
-
AIC <(aic-options)>
-
displays Akaike’s information criterion (AIC) for the models examined when you request variable selection with the SELECTION=
option in the MODEL
statement.
The following aic-options are available for models where you request the RSQUARE, ADJRSQ, or CP selection method:
-
LABEL
-
requests that the model number corresponding to the one displayed in the "Subset Selection Summary" table be used to label
the model with the smallest AIC statistic at each value of the number of parameters.
-
LABELVARS
-
requests that the list (excluding the intercept) of the regressors in the relevant model be used to label the model with the
smallest AIC statistic at each value of the number of parameters.
-
ALL
-
produces all appropriate plots.
-
BIC <(bic-options)>
-
displays Sawa’s Bayesian information criterion (BIC) for the models examined when you request variable selection with the
SELECTION= option in the MODEL
statement.
The following bic-options are available for models where you request the RSQUARE, ADJRSQ, or CP selection method:
-
LABEL
-
requests that the model number corresponding to the one displayed in the "Subset Selection Summary" table be used to label
the model with the smallest BIC statistic at each value of the number of parameters.
-
LABELVARS
-
requests that the list (excluding the intercept) of the regressors in the relevant model be used to label the model with the
smallest BIC statistic at each value of the number of parameters.
-
COOKSD <(LABEL)>
-
plots Cook’s D statistic by observation number. Observations whose Cook’s D statistic lies above the horizontal reference line at value 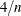 , where n is the number of observations used, are deemed to be influential (Rawlings, Pantula, and Dickey, 1998). If you specify the LABEL option, then points deemed as influential are labeled. If you do not specify an ID variable, the
observation number within the current BY group is used as the label. If you specify one or more ID variables in one or more
ID statements, then the first ID variable you specify is used for the labeling.
, where n is the number of observations used, are deemed to be influential (Rawlings, Pantula, and Dickey, 1998). If you specify the LABEL option, then points deemed as influential are labeled. If you do not specify an ID variable, the
observation number within the current BY group is used as the label. If you specify one or more ID variables in one or more
ID statements, then the first ID variable you specify is used for the labeling.
-
CP <(cp-options)>
-
displays Mallows’ 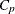 statistic for the models examined when you request variable selection with the SELECTION= option in the MODEL
statement. For models where you request the RSQUARE, ADJRSQ, or CP selection, reference lines corresponding to the equations
statistic for the models examined when you request variable selection with the SELECTION= option in the MODEL
statement. For models where you request the RSQUARE, ADJRSQ, or CP selection, reference lines corresponding to the equations
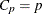 and
and 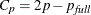 , where
, where 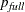 is the number of parameters in the full model (excluding the intercept) and p is the number of parameters in the subset model (including the intercept), are displayed on the plot of
is the number of parameters in the full model (excluding the intercept) and p is the number of parameters in the subset model (including the intercept), are displayed on the plot of 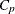 versus p. For the purpose of parameter estimation, Hocking (1976) suggests selecting a model where
versus p. For the purpose of parameter estimation, Hocking (1976) suggests selecting a model where 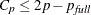 . For the purpose of prediction, Hocking suggests the criterion
. For the purpose of prediction, Hocking suggests the criterion 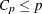 . Mallows (1973) suggests that all subset models with
. Mallows (1973) suggests that all subset models with 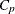 small and near p be considered for further study.
small and near p be considered for further study.
The following cp-options are available for models where you request the RSQUARE, ADJRSQ, or CP selection method:
-
LABEL
-
requests that the model number corresponding to the one displayed in the "Subset Selection Summary" table be used to label
the model with the smallest 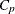 statistic at each value of the number of parameters.
statistic at each value of the number of parameters.
-
LABELVARS
-
requests that the list (excluding the intercept) of the regressors in the relevant model be used to label the model with the
smallest 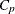 statistic at each value of the number of parameters.
statistic at each value of the number of parameters.
-
CRITERIA | CRITERIONPANEL <(criteria-options)>
-
produces a panel of fit criteria for the models examined when you request variable selection with the SELECTION= option in
the MODEL
statement. The fit criteria displayed are R-square, adjusted R-square, Mallows’ 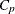 , Akaike’s information criterion (AIC), Sawa’s Bayesian information criterion (BIC), and Schwarz’s Bayesian information criterion
(SBC). For SELECTION=RSQUARE, SELECTION=ADJRSQ, or SELECTION=CP, scatter plots of these statistics versus the number of parameters
(including the intercept) are displayed. For other selection methods, line plots of these statistics as function of the selection
step number are displayed.
, Akaike’s information criterion (AIC), Sawa’s Bayesian information criterion (BIC), and Schwarz’s Bayesian information criterion
(SBC). For SELECTION=RSQUARE, SELECTION=ADJRSQ, or SELECTION=CP, scatter plots of these statistics versus the number of parameters
(including the intercept) are displayed. For other selection methods, line plots of these statistics as function of the selection
step number are displayed.
The following criteria-options are available:
-
LABEL
-
requests that the model number corresponding to the one displayed in the "Subset Selection Summary" table be used to label
the best model at each value of the number of parameters. This option applies only to the RSQUARE, ADJRSQ, and CP selection
methods.
-
LABELVARS
-
requests that the list (excluding the intercept) of the regressors in the relevant model be used to label the best model at
each value of the number of parameters. Since these labels are typically long, LABELVARS is supported only when the panel
is unpacked. This option applies only to the RSQUARE, ADJRSQ, and CP selection methods.
-
UNPACK
-
suppresses paneling. Separate plots are produced for each of the six fit statistics. For models where you request the RSQUARE,
ADJRSQ, or CP selection, two reference lines corresponding to the equations 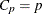 and
and 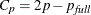 , where
, where 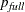 is the number of parameters in the full model (excluding the intercept) and p is the number of parameters in the subset model (including the intercept), are displayed on the plot of
is the number of parameters in the full model (excluding the intercept) and p is the number of parameters in the subset model (including the intercept), are displayed on the plot of 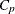 versus p. For the purpose of parameter estimation, Hocking (1976) suggests selecting a model where
versus p. For the purpose of parameter estimation, Hocking (1976) suggests selecting a model where 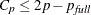 . For the purpose of prediction, Hocking suggests the criterion
. For the purpose of prediction, Hocking suggests the criterion 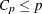 . Mallows (1973) suggests that all subset models with
. Mallows (1973) suggests that all subset models with 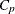 small and near p be considered for further study.
small and near p be considered for further study.
-
DFBETAS <(DFBETAS-options)>
-
produces panels of DFBETAS by observation number for the regressors in the model. Note that each panel contains at most six
plots, and multiple panels are used in the case where there are more than six regressors (including the intercept) in the
model. Observations whose DFBETAS’ statistics for a regressor are greater in magnitude than 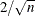 , where n is the number of observations used, are deemed to be influential for that regressor (Rawlings, Pantula, and Dickey, 1998).
, where n is the number of observations used, are deemed to be influential for that regressor (Rawlings, Pantula, and Dickey, 1998).
The following DFBETAS-options are available:
-
COMMONAXES
-
specifies that the same DFBETAS axis be used in all panels when multiple panels are needed. By default, the DFBETAS axis is
chosen independently for each panel. If you also specify the UNPACK option, then the same DFBETAS axis is used for each regressor.
-
LABEL
-
specifies that observations whose magnitude are greater than 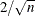 be labeled. If you do not specify an ID variable, the observation number within the current BY group is used as the label.
If you specify one or more ID variables on one or more ID statements, then the first ID variable you specify is used for the
labeling.
be labeled. If you do not specify an ID variable, the observation number within the current BY group is used as the label.
If you specify one or more ID variables on one or more ID statements, then the first ID variable you specify is used for the
labeling.
-
UNPACK
-
suppresses paneling. The DFBETAS statistics for each regressor are displayed on separate plots.
-
DFFITS <(LABEL)>
-
plots the DFFITS statistic by observation number. Observations whose DFFITS’ statistic is greater in magnitude than 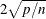 , where n is the number of observations used and p is the number of regressors, are deemed to be influential (Rawlings, Pantula, and Dickey, 1998). If you specify the LABEL option, then these influential observations are labeled. If you do not specify an ID variable,
the observation number within the current BY group is used as the label. If you specify one or more ID variables in one or
more ID statements, then the first ID variable you specify is used for the labeling.
, where n is the number of observations used and p is the number of regressors, are deemed to be influential (Rawlings, Pantula, and Dickey, 1998). If you specify the LABEL option, then these influential observations are labeled. If you do not specify an ID variable,
the observation number within the current BY group is used as the label. If you specify one or more ID variables in one or
more ID statements, then the first ID variable you specify is used for the labeling.
-
DIAGNOSTICS <(diagnostics-options)>
-
produces a summary panel of fit diagnostics:
-
residuals versus the predicted values
-
studentized residuals versus the predicted values
-
studentized residuals versus the leverage
-
normal quantile plot of the residuals
-
dependent variable values versus the predicted values
-
Cook’s D versus observation number
-
histogram of the residuals
-
"Residual-Fit" (or RF) plot consisting of side-by-side quantile plots of the centered fit and the residuals
-
box plot of the residuals if you specify the STATS=NONE suboption
You can specify the following diagnostics-options:
-
STATS=stats-options
-
determines which model fit statistics are included in the panel. See the global STATS= suboption for details. The PLOTS= suboption
of the DIAGNOSTICSPANEL option overrides the global PLOTS= suboption.
-
UNPACK
-
produces the eight plots in the panel as individual plots. Note that you can also request individual plots in the panel by
name without having to unpack the panel.
-
FITPLOT | FIT <(fit-options)>
-
produces a scatter plot of the data overlaid with the regression line, confidence band, and prediction band for models that
depend on at most one regressor excluding the intercept. When the number of points exceeds the MAXPOINTS=max value, a heat map is displayed instead of a scatter plot. By default, heat maps are not displayed if the number of observations
times the number of independent variables is greater than 150,000. See the MAXPOINTS=
option.
You can specify the following fit-options:
-
NOCLI
-
suppresses the prediction limits.
-
NOCLM
-
suppresses the confidence limits.
-
NOLIMITS
-
suppresses the confidence and prediction limits.
-
STATS=stats-options
-
determines which model fit statistics are included in the panel. See the global STATS= suboption for details. The PLOTS= suboption
of the FITPLOT option overrides the global PLOTS= suboption.
-
OBSERVEDBYPREDICTED <(LABEL)>
-
plots dependent variable values by the predicted values. If you specify the LABEL option, then points deemed as outliers or
influential (see the RSTUDENTBYLEVERAGE option for details) are labeled.
-
NONE
-
suppresses all plots.
-
PARTIAL <(UNPACK)>
-
produces panels of partial regression plots for each regressor with at most six regressors per panel. If you specify the UNPACK
option, then all partial plot panels are unpacked.
-
PREDICTIONS (X=numeric-variable <prediction-options>)
-
produces a panel of two plots whose horizontal axis is the variable you specify in the required X= suboption. The upper plot
in the panel is a scatter plot of the residuals. The lower plot shows the data overlaid with the regression line, confidence
band, and prediction band. This plot is appropriate for models where all regressors are known to be functions of the single
variable that you specify in the X= suboption.
You can specify the following prediction-options:
-
NOCLI
-
suppresses the prediction limits.
-
NOCLM
-
suppresses the confidence limits
-
NOLIMITS
-
suppresses the confidence and prediction limits
-
SMOOTH
-
requests a nonparametric smoothing of the residuals as a function of the variable you specify in the X= suboption. This nonparametric
fit is a loess fit that uses local linear polynomials, linear interpolation, and a smoothing parameter that is selected to
yield a local minimum of the corrected Akaike’s information criterion (AICC). See Chapter 59: The LOESS Procedure, for details. The SMOOTH option is not supported when a FREQ
statement is used.
-
UNPACK
-
suppresses paneling.
-
QQPLOT | QQ
-
produces a normal quantile plot of the residuals.
-
RESIDUALBOXPLOT | BOXPLOT <(LABEL)>
-
produces a box plot consisting of the residuals. If you specify label option, points deemed far-outliers are labeled. If you
do not specify an ID variable, the observation number within the current BY group is used as the label. If you specify one
or more ID variables in one or more ID statements, then the first ID variable you specify is used for the labeling.
-
RESIDUALBYPREDICTED <(LABEL)>
-
plots residuals by predicted values. If you specify the LABEL option, then points deemed as outliers or influential (see the
RSTUDENTBYLEVERAGE option for details) are labeled.
-
RESIDUALCHART <(residual-chart-options)>
RC <(residual-chart-options)>
-
produces the residual chart and enables you to specify residual-chart-options. This chart displays studentized residuals and Cook’s D in side-by-side bar charts. This chart is also displayed when you specify the R option in the MODEL statement.
Unlike most graphs, the height of this chart can vary as a function of the number of observations that appear in the chart.
You can specify the following residual-chart-options to control the height and other aspects of the chart:
-
COMPUTEHEIGHT=a b <max>
CH=a b <max>
-
specifies the constants for computing the height of the chart. For n dimensions, intercept a, slope b, and maximum height max, the height is min(a + b (n + 1), max). By default, COMPUTEHEIGHT=150 15 1650. Thus, the default height in pixels is min(150 + 15(n + 1), 1650). The default unit is pixels, and you can use the UNIT= residual-chart-option to change the unit to inches or centimeters.
-
MAX=max
-
species the maximum number of points to display in each chart. When the number of points exceeds max, charts of up to max observations are displayed until all observations are displayed.
-
SETHEIGHT=height
SH=height
-
specifies the height of the chart. By default, the height is based on the COMPUTEHEIGHT= option. The default unit is pixels,
and you can use the UNIT= residual-chart-option to change the unit to inches or centimeters.
-
UNIT=PX | IN | CM
-
specifies the unit (pixels, inches, or centimeters) for the SETHEIGHT= and COMPUTEHEIGHT= residual-chart-options. Inches equals pixels divided by 96, and centimeters equals inches times 2.54. By default, UNIT=PX.
-
UNPACK
-
suppresses paneling. The studentized residuals and Cook’s D are displayed in separate charts. When you specify the UNPACK residual-chart-option, residuals, standard errors, and other values that go into the computations are added to each chart.
-
RESIDUALS <(residual-options)>
-
produces panels of the residuals versus the regressors in the model. Each panel contains at most six plots, and multiple panels
are used when the model contains more than six regressors (including the intercept). When the number of points exceeds the
MAXPOINTS=max value, a heat map is displayed instead of a scatter plot. By default, heat maps are not displayed if the number of observations
times the number of independent variables is greater than 150,000. See the MAXPOINTS=
option. You can specify the following residual-options:
-
SMOOTH
-
requests a nonparametric smoothing of the residuals for each regressor. Each nonparametric fit is a loess fit that uses local
linear polynomials, linear interpolation, and a smoothing parameter that is selected to yield a local minimum of the corrected
Akaike’s information criterion (AICC). See Chapter 59: The LOESS Procedure, for details. The SMOOTH option is not supported when a FREQ
statement is used.
-
UNPACK
-
suppresses paneling.
-
RESIDUALHISTOGRAM
-
produces a histogram of the residuals.
-
RFPLOT | RF
-
produces a "Residual-Fit" (or RF) plot consisting of side-by-side quantile plots of the centered fit and the residuals. This
plot "shows how much variation in the data is explained by the fit and how much remains in the residuals" (Cleveland, 1993).
-
RIDGE | RIDGEPANEL | RIDGEPLOT <(ridge-options)>
-
creates panels of VIF values and standardized ridge estimates by ridge values for each coefficient. The VIF values for each
coefficient are connected by lines and are displayed in the upper plot in each panel. The points corresponding to the standardized
estimates of each coefficient are connected by lines and are displayed in the lower plot in each panel. By default, at most
10 coefficients are represented in a panel and multiple panels are produced for models with more than 10 regressors. For ridge
estimates to be computed and plotted, the OUTEST= option must be specified in the PROC REG
statement, and the RIDGE= list must be specified in either the PROC REG
or the MODEL
statement. (See Example 85.5.)
The following ridge-options are available:
-
COMMONAXES
-
specifies that the same VIF axis and the same standardized estimate axis are used in all panels when multiple panels are needed.
By default, these axes are chosen independently for the regressors shown in each panel.
-
RIDGEAXIS=LINEAR | LOG
-
specifies the axis type used to display the ridge parameters. The default is RIDGEAXIS=LINEAR. Note that the point with the
ridge parameter equal to zero is not displayed if you specify RIDGEAXIS=LOG.
-
UNPACK
-
suppresses paneling. The traces of the VIF statistics and standardized estimates are shown in separate plots.
-
VARSPERPLOT=ALL
VARSPERPLOT=number
-
specifies the maximum number of regressors displayed in each panel or in each plot if you additionally specify the UNPACK
option. If you specify VARSPERPLOT=ALL, then the VIF values and ridge traces for all regressors are displayed in a single
panel.
-
VIFAXIS=LINEAR | LOG
-
specifies the axis type used to display the VIF statistics. The default is VIFAXIS=LINEAR.
-
RSQUARE <(rsquare-options)>
-
displays the R-square values for the models examined when you request variable selection with the SELECTION= option in the
MODEL
statement.
The following rsquare-options are available for models where you request the RSQUARE, ADJRSQ, or CP selection method:
-
LABEL
-
requests that the model number corresponding to the one displayed in the "Subset Selection Summary" table be used to label
the model with the largest R-square statistic at each value of the number of parameters.
-
LABELVARS
-
requests that the list (excluding the intercept) of the regressors in the relevant model be used to label the model with the
largest R-square statistic at each value of the number of parameters.
-
RSTUDENTBYLEVERAGE <(LABEL)>
-
plots studentized residuals by leverage. Observations whose studentized residuals lie outside the band between the reference
lines 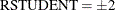 are deemed outliers. Observations whose leverage values are greater than the vertical reference
are deemed outliers. Observations whose leverage values are greater than the vertical reference 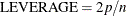 , where p is the number of parameters including the intercept and n is the number of observations used, are deemed influential (Rawlings, Pantula, and Dickey, 1998). If you specify the LABEL option, then points deemed as outliers or influential are labeled. If you do not specify an ID
variable, the observation number within the current BY group is used as the label. If you specify one or more ID variables
in one or more ID statements, then the first ID variable you specify is used for the labeling.
, where p is the number of parameters including the intercept and n is the number of observations used, are deemed influential (Rawlings, Pantula, and Dickey, 1998). If you specify the LABEL option, then points deemed as outliers or influential are labeled. If you do not specify an ID
variable, the observation number within the current BY group is used as the label. If you specify one or more ID variables
in one or more ID statements, then the first ID variable you specify is used for the labeling.
-
RSTUDENTBYPREDICTED <(LABEL)>
-
plots studentized residuals by predicted values. If you specify the LABEL option, then points deemed as outliers or influential
(see the RSTUDENTBYLEVERAGE option for details) are labeled.
-
SBC <(sbc-options)>
-
displays Schwarz’s Bayesian information criterion (SBC) for the models examined when you request variable selection with the
SELECTION= option in the MODEL
statement.
The following sbc-options are available for models where you request the RSQUARE, ADJRSQ, or CP selection method:
-
LABEL
-
requests that the model number corresponding to the one displayed in the "Subset Selection Summary" table be used to label
the model with the smallest SBC statistic at each value of the number of parameters.
-
LABELVARS
-
requests that the list (excluding the intercept) of the regressors in the relevant model be used to label the model with the
smallest SBC statistic at each value of the number of parameters.


 Missing ValuesInput Data SetsOutput Data SetsInteractive AnalysisModel-Selection MethodsCriteria Used in Model-Selection MethodsLimitations in Model-Selection MethodsParameter Estimates and Associated StatisticsPredicted and Residual ValuesModels of Less Than Full RankCollinearity DiagnosticsModel Fit and Diagnostic StatisticsInfluence StatisticsReweighting Observations in an AnalysisTesting for HeteroscedasticityTesting for Lack of FitMultivariate TestsAutocorrelation in Time Series DataComputations for Ridge Regression and IPC AnalysisConstruction of Q-Q and P-P PlotsComputational MethodsComputer Resources in Regression AnalysisDisplayed OutputPlot Options Superseded by ODS GraphicsODS Table NamesODS Graphics
Missing ValuesInput Data SetsOutput Data SetsInteractive AnalysisModel-Selection MethodsCriteria Used in Model-Selection MethodsLimitations in Model-Selection MethodsParameter Estimates and Associated StatisticsPredicted and Residual ValuesModels of Less Than Full RankCollinearity DiagnosticsModel Fit and Diagnostic StatisticsInfluence StatisticsReweighting Observations in an AnalysisTesting for HeteroscedasticityTesting for Lack of FitMultivariate TestsAutocorrelation in Time Series DataComputations for Ridge Regression and IPC AnalysisConstruction of Q-Q and P-P PlotsComputational MethodsComputer Resources in Regression AnalysisDisplayed OutputPlot Options Superseded by ODS GraphicsODS Table NamesODS Graphics