The FASTCLUS Procedure
Computational Resources
Let

Memory
The memory required is approximately 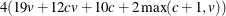 bytes.
bytes.
If you request the DISTANCE option, an additional 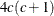 bytes of space is needed.
bytes of space is needed.
Time
The overall time required by PROC FASTCLUS is roughly proportional to 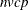 if c is small with respect to n.
if c is small with respect to n.
Initial seed selection requires one pass over the data set. If the observations are in random order, the time required is roughly proportional to
![\[ nvc + vc^2 \]](images/statug_fastclus0032.png)
unless you specify REPLACE=NONE. In that case, a complete pass might not be necessary, and the time is roughly proportional
to 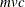 , where
, where 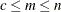 .
.
The DRIFT option, each iteration, and the final assignment of cluster seeds each require one pass, with time for each pass
roughly proportional to 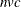 .
.
For greatest efficiency, you should list the variables in the VAR statement in order of decreasing variance.
