-
ABSENT=number | qs
-
specifies the value to be used as an absence value in an irrelevant absent-absent match for all of the asymmetric nominal variables. If you want to specify a different absence value for a particular variable, use the
ABSENT= option in the VAR statement. See the ABSENT= option in the section VAR Statement for details.
An absence value for a variable can be either a numeric value or a quoted string consisting of combinations of characters.
For instance, ., -999, and "NA" are legal values for the ABSENT= option.
The default absence value for a character variable is "NONE" (notice that a blank value is considered a missing value), and
the default absence value for a numeric variable is 0.
-
ADD=c
-
specifies a constant, c, to add to each value after standardizing and multiplying by the value you specify in the MULT= option. The default value is 0.
-
DATA=SAS-data-set
-
specifies the input data set containing observations from which the proximity is computed. If you omit the DATA= option, the most recently created SAS data set is used.
-
FUZZ=c
-
specifies the relative fuzz factor for computing the standardized scores. The default value is 1E–14. For the OUTSDZ= data set, the score is computed as follows:
where m is the numeric constant specified in the MULT= option, or 1 if MULT= option is not specified.
-
INITIAL=method
-
specifies the method of computing initial estimates for the A-estimates (ABW, AWAVE, and AHUBER). The following methods are not allowed for the INITIAL= option: ABW, AHUBER, AWAVE, and
IN.
The default value is INITIAL=MAD.
-
METHOD=method
-
specifies the method of computing proximity measures.
For use in PROC CLUSTER, distance or dissimilarity measures such as METHOD=EUCLID or METHOD=DGOWER should be chosen.
The following six tables outline the proximity measures available for the METHOD= option. These tables are classified by levels
of measurement accepted by each method. Each table contains four or five columns: the Method column shows the proximity measures,
one or two Range columns show the upper and lower bounds, and the TYPE= column shows the type of proximity. The TYPE= column
contains SIMILAR if a method generates similarity measures or DISTANCE if a method generates distance or dissimilarity measures.
The output data set is of the type shown. For more information about the output data set, see the OUT=
option.
For formulas and descriptions of these methods, see the section Details: DISTANCE Procedure.
Table 36.2 shows the range and output matrix type of the GOWER and DGOWER methods. These two methods accept all measurement levels including
ratio, interval, ordinal, nominal, and asymmetric nominal. METHOD=GOWER or METHOD=DGOWER always implies standardization. Assuming
all the numeric (ordinal, interval, and ratio) variables are standardized by their corresponding default methods, the possible
range values for both methods are from 0 and 1, inclusive. For more information about the default methods of standardization
for METHOD=GOWER or METHOD=DGOWER, see the STD= option in the section VAR Statement.
Table 36.2: Methods That Accept All Measurement Levels
|
Method
|
Description
|
Range
|
TYPE=
|
|
GOWER
|
Gower and Legendre (1986) similarity
|
0 to 1
|
SIMILAR
|
|
DGOWER
|
1 minus GOWER
|
0 to 1
|
DISTANCE
|
Table 36.3 shows methods that accept ratio, interval, and ordinal variables.
Table 36.3: Methods That Accept Ratio, Interval, and Ordinal Variables
|
Method
|
Description
|
Range
|
TYPE=
|
|
EUCLID
|
Euclidean distance
|
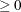
|
DISTANCE
|
|
SQEUCLID
|
Squared Euclidean distance
|
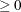
|
DISTANCE
|
|
SIZE
|
Size distance
|
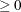
|
DISTANCE
|
|
SHAPE
|
Shape distance
|
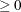
|
DISTANCE
|
|
COV
|
Covariance
|
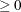
|
SIMILAR
|
|
CORR
|
Correlation
|
–1 to 1
|
SIMILAR
|
|
DCORR
|
Correlation transformed to Euclidean distance
|
0 to 2
|
DISTANCE
|
|
SQCORR
|
Squared correlation
|
0 to 1
|
SIMILAR
|
|
DSQCORR
|
One minus squared correlation
|
0 to 1
|
DISTANCE
|
|
L(p)
|
Minkowski (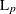 ) distance, where p is a positive numeric value ) distance, where p is a positive numeric value
|
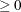
|
DISTANCE
|
|
CITYBLOCK
|
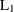 , city-block, or Manhattan distance , city-block, or Manhattan distance
|
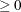
|
DISTANCE
|
|
CHEBYCHEV
|
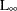
|
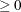
|
DISTANCE
|
|
POWER(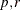 ) )
|
Generalized Euclidean distance where p is a positive numeric value and r is a nonnegative numeric value. The distance between two observations is the rth root of sum of the absolute differences to the pth power between the values for the observations.
|
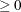
|
DISTANCE
|
Table 36.4 shows methods that accept ratio variables. Notice that all possible range values are nonnegative, because ratio variables are assumed to
be positive.
Table 36.4: Methods That Accept Ratio Variables
|
Method
|
Description
|
Range
|
TYPE=
|
|
SIMRATIO
|
Similarity ratio (if variables are binary, this is the Jaccard coefficient)
|
0 to 1
|
SIMILAR
|
|
DISRATIO
|
One minus similarity ratio
|
0 to 1
|
DISTANCE
|
|
NONMETRIC
|
Lance and Williams nonmetric coefficient
|
0 to 1
|
DISTANCE
|
|
CANBERRA
|
Canberra metric distance coefficient
|
0 to 1
|
DISTANCE
|
|
COSINE
|
Cosine coefficient
|
0 to 1
|
SIMILAR
|
|
DOT
|
Dot (inner) product coefficient
|
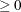
|
SIMILAR
|
|
OVERLAP
|
Overlap similarity
|
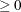
|
SIMILAR
|
|
DOVERLAP
|
Overlap dissimilarity
|
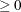
|
DISTANCE
|
|
CHISQ
|
Chi-square coefficient
|
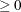
|
DISTANCE
|
|
CHI
|
Square root of chi-square coefficient
|
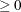
|
DISTANCE
|
|
PHISQ
|
Phi-square coefficient
|
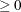
|
DISTANCE
|
|
PHI
|
Square root of phi-square coefficient
|
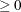
|
DISTANCE
|
Table 36.5 shows methods that accept nominal variables.
Table 36.5: Methods That Accept Nominal Variables
|
Method
|
Description
|
Range
|
TYPE=
|
|
HAMMING
|
Hamming distance
|
0 to v
|
DISTANCE
|
|
MATCH
|
Simple matching coefficient
|
0 to 1
|
SIMILAR
|
|
DMATCH
|
Simple matching coefficient transformed to Euclidean distance
|
0 to 1
|
DISTANCE
|
|
DSQMATCH
|
Simple matching coefficient transformed to squared Euclidean distance
|
0 to 1
|
DISTANCE
|
|
HAMANN
|
Hamann coefficient
|
–1 to 1
|
SIMILAR
|
|
RT
|
Roger and Tanimoto
|
0 to 1
|
SIMILAR
|
|
SS1
|
Sokal and Sneath 1
|
0 to 1
|
SIMILAR
|
|
SS3
|
Sokal and Sneath 3
|
0 to 1
|
SIMILAR
|
Note that v denotes the number of variables (dimensionality).
Table 36.6 shows methods that accept asymmetric nominal variables. Use the ABSENT= option to create a value to be considered absent.
Table 36.6: Methods That Accept Asymmetric Nominal Variables
|
Method
|
Description
|
Range
|
TYPE=
|
|
DICE
|
Dice coefficient or Czekanowski/Sorensen similarity coefficient
|
0 to 1
|
SIMILAR
|
|
RR
|
Russell and Rao
|
0 to 1
|
SIMILAR
|
|
BLWNM
|
Binary Lance and Williams nonmetric, or Bray-Curtis coefficient
|
0 to 1
|
DISTANCE
|
|
K1
|
Kulcynski 1
|
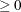
|
SIMILAR
|
Table 36.7 shows methods that accept asymmetric nominal and ratio variables. Use the ABSENT= option to create a value to be considered absent. The
table contains five columns. The third column contains possible range values if only one level of measurement (either ratio
or asymmetric nominal but not both) is specified; the fourth column contains possible range values if both levels are specified.
The JACCARD method is equivalent to the SIMRATIO method if there is no asymmetric nominal variable; if both ratio and asymmetric
nominal variables are present, the coefficient is computed as the sum of the coefficient from the ratio variables and the
coefficient from the asymmetric nominal variables. See "Proximity Measures" in the section Details: DISTANCE Procedure for the formula and descriptions of the JACCARD method.
Table 36.7: Methods That Accept Asymmetric Nominal and Ratio Variables
|
Method
|
Description
|
Range (One Level)
|
Range (Two Levels)
|
TYPE=
|
|
JACCARD
|
Jaccard similarity
coefficient
|
0 to 1
|
0 to 2
|
SIMILAR
|
|
DJACCARD
|
Jaccard dissimilarity
coefficient
|
0 to 1
|
0 to 2
|
DISTANCE
|
-
MULT=c
-
specifies a numeric constant, c, by which to multiply each value after standardizing. The default value is 1.
-
NOMISS
-
omits observations with missing values from computation of the location and scale measures when standardizing; generates undefined (missing) distances for observations
with missing values when computing distances. Use the UNDEF= option to specify the undefined values.
If a distance matrix is created to be used as an input to PROC CLUSTER, the NOMISS option should not be used because PROC
CLUSTER does not accept distance matrices with missing values.
-
NORM
-
normalizes the scale estimator to be consistent for the standard deviation of a normal distribution when you specify the option STD=AGK, STD=IQR, STD=MAD, or STD=SPACING in the VAR statement.
-
NOSTD
-
suppresses standardization of the variables. The NOSTD option should not be specified with the STDONLY option or with the REPLACE option.
-
OUT=SAS-data-set
-
specifies the name of the SAS data set created by PROC DISTANCE. The output data set contains the BY variables, the ID variable, computed distance variables, the COPY variables, the FREQ
variable, and the WEIGHT variables.
If you omit the OUT= option, PROC DISTANCE creates an output data set named according to the DATAn convention.
The output data set is of type TYPE=DISTANCE or TYPE=SIMILAR. See the METHOD=
option for more information about the association between the method and the output data set type. Data set types do not
persist when you copy or modify a data set. You must specify the TYPE= data set option for the new data set, as in the following
example:
data dist2(type=distance);
set dist;
run;
If you do not specify the TYPE=DISTANCE data set option, the new data set is the default TYPE=DATA. If you use the new data
set in a procedure that accepts both TYPE=DATA or TYPE=DISTANCE data sets (such as PROC CLUSTER or PROC MODECLUS), the results
will be incorrect.
-
OUTSDZ=SAS-data-set
-
specifies the name of the SAS data set containing the standardized scores. The output data set contains a copy of the DATA= data set, except that the analyzed variables have been standardized.
Analyzed variables are those listed in the VAR statement.
-
PREFIX=name
-
specifies a prefix for naming the distance variables in the OUT= data set. By default, the names are Dist1, Dist2, …, Distn. If you specify PREFIX=ABC, the variables are named ABC1, ABC2, …, ABCn. If the ID statement is also specified, the variables are named by appending the value of the ID variable to the prefix.
-
RANKSCORE=MIDRANK | INDEX
-
specifies the method of assigning scores to ordinal variables. The available methods are listed as follows:
- MIDRANK
-
assigns consecutive integers to each category with consideration of the frequency value. This is the default method.
- INDEX
-
assigns consecutive integers to each category regardless of frequencies.
The following example explains how each method assigns the rank scores. Suppose the data contain an ordinal variable ABC with values A, B, C. There are two ways to assign numbers. One is to use midranks, which depend on the frequencies of each
category. Another is to assign consecutive integers to each category, regardless of frequencies.
Table 36.8: Example of Assigning Rank Scores
|
ABC
|
MIDRANK
|
INDEX
|
|
A
|
1.5
|
1
|
|
A
|
1.5
|
1
|
|
B
|
4
|
2
|
|
B
|
4
|
2
|
|
B
|
4
|
2
|
|
C
|
6
|
3
|
-
REPLACE
-
replaces missing data with zero in the standardized data (to correspond to the location measure before standardizing). To replace missing data with something else,
use the MISSING= option in the VAR statement. The REPLACE option implies standardization.
You cannot specify the following options together:
-
REPONLY
-
measure specified by the MISSING= option or the STD= option (if the MISSING= option is not specified), but does not standardize the data. If the MISSING= option is not specified and METHOD=GOWER is specified, missing values are replaced
by the location measure from the RANGE method (the minimum value), no matter what the value of the STD= option is.
You cannot specify both the REPLACE and the REPONLY options.
-
SHAPE=TRIANGLE | TRI | SQUARE | SQU | SQR
-
specifies the shape of the proximity matrix to be stored in the OUT= data set. SHAPE=TRIANGLE requests the matrix to be stored as a lower triangular matrix; SHAPE=SQUARE requests that the
matrix be stored as a square matrix. Use SHAPE=SQUARE if the output data set is to be used as input to the MODECLUS procedures.
The default is TRIANGLE.
-
SNORM
-
normalizes the scale estimator to have an expectation of approximately 1 for a standard normal distribution when the STD=SPACING option is specified.
-
STDONLY
-
standardizes variables only and computes no distance matrix. You must use the OUTSDZ= option to save the standardized scores. You cannot specify both the STDONLY option and the
NOSTD option.
-
UNDEF=n
-
specifies the numeric constant used to replace undefined distances, such as when an observation has all missing values, or if a divisor is zero.
-
VARDEF=DF | N | WDF | WEIGHT | WGT
-
specifies the divisor to be used in the calculation of distance, dissimilarity, or similarity measures, and for standardizing variables whenever a variance or covariance is computed.
By default, VARDEF=DF. The values and associated divisors are as follows:
|
Value
|
Divisor
|
Formula
|
|
DF
|
degrees of freedom
|
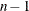
|
|
N
|
number of observations
|
n
|
|
WDF
|
sum of weights minus 1
|
(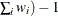
|
|
WEIGHT | WGT
|
sum of weights
|
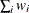
|
