| The MODECLUS Procedure |
| PROC MODECLUS Statement |
The PROC MODECLUS statement starts the MODECLUS procedure. The options available with the PROC MODECLUS statement are summarized in Table 57.1 and discussed in the following sections.
Option |
Description |
|---|---|
Specify input and output data sets |
|
specifies input data set name |
|
specifies output data set name for observations |
|
specifies output data set name for clusters |
|
specifies output data set name for cluster solutions |
|
Specify variables in output data sets |
|
specifies variable in the OUT= and OUTCLUS= data sets identifying clusters |
|
specifies variable in the OUT= data set containing density estimates |
|
specifies length of variables in the output data sets |
|
Summarize and process coordinate data before clustering |
|
requests simple statistics |
|
standardizes the variables to mean 0 and standard deviation 1 |
|
Specify smoothing parameters |
|
specifies number of neighbors to use for |
|
specifies number of neighbors to use for clustering |
|
specifies number of neighbors to use for |
|
specifies radius of the sphere of support for uniform-kernel density estimation |
|
specifies radius of the neighborhood for clustering |
|
specifies radius of the sphere of support for uniform-kernel density estimation and the neighborhood clustering |
|
Specify density estimation options |
|
specifies number of times the density estimates are to be cascaded |
|
CROSS or CROSSLIST |
computes the likelihood cross validation criterion |
specifies dimensionality to be used when computing density estimates |
|
uses arithmetic means for cascading density estimates |
|
uses harmonic means for cascading density estimates |
|
uses sums for cascading density estimates |
|
Specify clustering methods and options |
|
dissolves clusters with n or fewer members |
|
stops the analysis after obtaining a solution with either no cluster or a single cluster |
|
requests that nonsignificant clusters be hierarchically joined |
|
specifies maximum number of clusters to be obtained with METHOD=6 |
|
specifies clustering method to use |
|
specifies minimum members for either cluster to be designated a modal cluster when two clusters are joined using METHOD=5 |
|
specifies power of the density used with METHOD=6 |
|
specifies approximate significance tests for the number of clusters |
|
specifies assignment threshold used with METHOD=6 |
|
Specify the output display options |
|
produces all optional output |
|
displays the density and cluster membership of observations with neighbors belonging to a different cluster |
|
retains the neighbor lists for each observation in memory |
|
displays the estimated cross validated log density of each observation |
|
displays the estimated density and cluster membership of each observation |
|
displays estimates of local dimensionality and writes them to the OUT=data set |
|
displays the neighbors of each observation |
|
suppresses the display of the output |
|
suppresses the display of the summary of the number of clusters, number of unassigned observations, and maximum |
|
suppresses the display of statistics for each cluster |
|
traces the cluster assignments when METHOD=6 |
|
You can specify at least one of the following options for smoothing parameters for density estimation: DK=, K=, DR=, or R=. To obtain a cluster analysis, you can specify the METHOD= option and at least one of the following smoothing parameters for clustering: CK=, K=, CR=, or R=. If you want significance tests for the number of clusters, you should specify either the DR= or R= option. If none of the smoothing parameters is specified, the MODECLUS procedure provides a default value for the R= option. See the section Density Estimation for the formula of a reasonable first guess for R= and a discussion of smoothing parameters.
You can specify lists of values for the DK=, CK=, K=, DR=, CR=, and R= options. Numbers in the lists can be separated by blanks or commas. You can include in the lists one or more items of the form start TO stop BY increment. Each list can contain either one value or the same number of values as in every other list that contains more than one value. If a list has only one value, that value is used in combination with all the values in longer lists. If two or more lists have more than one value, then one analysis is done by using the first value in each list, another analysis is done by using the second value in each list, and so on.
You can specify the following options in the PROC MODECLUS statement.
- ALL
- AM
specifies arithmetic means for cascading density estimates. See the description of the CASCADE= option.
- BOUNDARY
displays the density and cluster membership of observations with neighbors belonging to a different cluster.
- CASCADE=n
- CASC=n
specifies the number of times the density estimates are to be cascaded (see the section Density Estimation). The default value 0 performs no cascading.
You can specify a list of values for the CASCADE= option. Each value in the list is combined with each combination of smoothing parameters to produce a separate analysis.
- CK=n
specifies the number of neighbors to use for clustering. The number of neighbors should be at least two but less than the number of observations. See the section Density Estimation for details.
- CLUSTER=name
provides a name for the variable in the OUT= and OUTCLUS= data sets identifying clusters. The default name is CLUSTER.
- CORE
keeps the neighbor lists for each observation in the computer memory to make small problems run faster.
- CR=n
specifies the radius of the neighborhood for clustering. See the section Density Estimation for details.
- CROSS
computes the likelihood cross validation criterion (Silverman; 1986, pp. 52–55). This option appears to be of limited usefulness. See the section Density Estimation for details.
- CROSSLIST
displays the cross validated log density of each observation.
- DATA=SAS-data-set
specifies the input data set containing observations to be clustered. If you omit the DATA= option, the most recently created SAS data set is used.
If the data set is TYPE=DISTANCE, the data are interpreted as a distance matrix. The number of variables must equal the number of observations in the data set or in each BY group. The distances are assumed to be Euclidean, but the procedure accepts other types of distances or dissimilarities. Unlike the CLUSTER procedure, PROC MODECLUS uses the entire distance matrix, not just the lower triangle; the distances are not required to be symmetric. The neighbors of a given observation are determined solely from the distances in that observation. Missing values are considered infinite. Various distance measures can be computed from coordinate data by using the DISTANCE procedure (for detailed information, see Chapter 32, The DISTANCE Procedure ).
If the data set is not TYPE=DISTANCE, the data are interpreted as coordinates in a Euclidean space, and Euclidean distances are computed. The variables can be discrete or continuous and should be at the interval level of measurement.
- DENSITY=name
provides a name for the variable in the OUT= data set containing density estimates. The default name is DENSITY.
- DIMENSION=n
- DIM=n
specifies the dimensionality to be used when computing density estimates. The default is the number of variables if the data are coordinates; the default is 1 if the data are distances.
- DK=n
specifies the number of neighbors to use for
 th-nearest-neighbor density estimation. The number of neighbors should be at least two but less than the number of observations. See the section Density Estimation for details.
th-nearest-neighbor density estimation. The number of neighbors should be at least two but less than the number of observations. See the section Density Estimation for details. - DOCK=n
dissolves clusters with n or fewer members by making the members unassigned.
- DR=n
specifies the radius of the sphere of support for uniform-kernel density estimation. See the section Density Estimation for details.
- EARLY
stops the cluster analysis after obtaining either a solution with no cluster or a solution with one cluster to which all observations are assigned. The smoothing parameters should be specified in increasing order. This can reduce the computer time required for the analysis but might occasionally miss some multiple-cluster solutions.
- HM
uses harmonic means for cascading density estimates. See the description of the CASCADE= option for details.
- JOIN<=p>
requests that nonsignificant clusters be hierarchically joined. The JOIN option implies the TEST option. After each solution is obtained, the cluster with the largest approximate
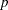 -value is either joined to a neighboring cluster or, if there is no neighboring cluster, dissolved by making all of its members unassigned. After two clusters are joined, an analysis of the remaining clusters is displayed.
-value is either joined to a neighboring cluster or, if there is no neighboring cluster, dissolved by making all of its members unassigned. After two clusters are joined, an analysis of the remaining clusters is displayed. If you do not specify a
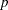 -value with the JOIN= option, joining continues until only one cluster remains, and the results are written to the output data sets after each analysis. If you specify a
-value with the JOIN= option, joining continues until only one cluster remains, and the results are written to the output data sets after each analysis. If you specify a 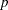 -value with the JOIN= option, joining continues until the greatest approximate
-value with the JOIN= option, joining continues until the greatest approximate 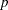 -value is less than the value given in the JOIN= option, and only if there is more than one cluster are the results for that analysis written to the output data sets.
-value is less than the value given in the JOIN= option, and only if there is more than one cluster are the results for that analysis written to the output data sets. Any value of
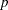 less than 1E
less than 1E 8 is set to 1E
8 is set to 1E 8.
8. - K=n
specifies the number of neighbors to use for
 th-nearest-neighbor density estimation and clustering. The number of neighbors should be at least two but less than the number of observations. Specifying K=n is equivalent to specifying both DK=n and CK=n. See the section Density Estimation for details.
th-nearest-neighbor density estimation and clustering. The number of neighbors should be at least two but less than the number of observations. Specifying K=n is equivalent to specifying both DK=n and CK=n. See the section Density Estimation for details. - LIST
displays the estimated density and cluster membership of each observation.
- LOCAL
requests estimates of local dimensionality (Tukey and Tukey; 1981, pp. 236–237).
- MAXCLUSTERS=n
- MAXC=n
specifies the maximum number of clusters to be obtained with the METHOD=6 option. By default, there is no fixed limit.
- METHOD=n
- MET=n
- M=n
specifies what clustering method to use. Since these methods do not have widely recognized names, the methods are indicated by numbers from 0 to 6. The methods are described in the section Clustering Methods. For most purposes, METHOD=1 is recommended, although METHOD=6 might occasionally produce better results in return for considerably greater computer time and space requirements. METHOD=1 is not good for discrete coordinate data with only a few equally spaced values. In this case, METHOD=6 or METHOD=3 works better. METHOD=4 or METHOD=5 is less desirable than other methods when there are ties, since a general characteristic of agglomerative hierarchical clustering methods is that the results are indeterminate in the presence of ties.
You must specify the METHOD= option to obtain a cluster analysis.
You can specify a list of values for the METHOD= option. Each value in the list is combined with each combination of smoothing and cascading parameters to produce a separate cluster analysis.
- MODE=n
specifies that when two clusters are joined using the METHOD=5 option (no other methods are affected by the MODE= option), each must have at least n members for either cluster to be designated a modal cluster. In any case, each cluster must also have a maximum density greater than the fusion density for either cluster to be designated a modal cluster. If you specify the K= option, the default value of the MODE= option is the same as the value of the K= option because the use of
 th-nearest-neighbor density estimation limits the resolution that can be obtained for clusters with fewer than
th-nearest-neighbor density estimation limits the resolution that can be obtained for clusters with fewer than  members. If you do not specify the K= option, the default is MODE=2. If you specify MODE=0, the default value is used instead of 0. If you specify a FREQ statement, the MODE= value is compared to the number of observations in each cluster, not to the sum of the frequencies.
members. If you do not specify the K= option, the default is MODE=2. If you specify MODE=0, the default value is used instead of 0. If you specify a FREQ statement, the MODE= value is compared to the number of observations in each cluster, not to the sum of the frequencies. - NEIGHBOR
displays the neighbors of each observation in a table called "Nearest Neighbor List." See Nearest Neighbor List for information displayed in the table.
- NOPRINT
suppresses the display of the output. Note that this option temporarily disables the Output Delivery System (ODS); see Chapter 20, Using the Output Delivery System, for more information.
- NOSUMMARY
suppresses the display of the summary of the number of clusters, number of unassigned observations, and maximum
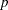 -value for each analysis.
-value for each analysis. - OUT=SAS-data-set
specifies the output data set containing the input data plus density estimates, cluster membership, and variables identifying the type of solution. There is an output observation corresponding to each input observation for each solution. Therefore, the OUT= data set can be very large.
If you want to create a permanent SAS data set, you must specify a two-level name. For details about OUT= data sets, see the section Output Data Sets. See SAS Language Reference: Concepts for more information about permanent SAS data sets.
- OUTCLUS=SAS-data-set
- OUTC=SAS-data-set
specifies the output data set containing an observation corresponding to each cluster in each solution. The variables identify the solution and contain statistics describing the clusters.
If you want to create a permanent SAS data set, you must specify a two-level name. For details about OUTCLUS= data sets, see the section Output Data Sets. See SAS Language Reference: Concepts for more information about permanent SAS data sets.
- OUTSUM=SAS-data-set
- OUTS=SAS-data-set
specifies the output data set containing an observation corresponding to each cluster solution, giving the number of clusters and the number of unclassified observations for that solution.
If you want to create a permanent SAS data set, you must specify a two-level name. For details about OUTSUM= data sets, see the section Output Data Sets. See SAS Language Reference: Concepts for more information about permanent SAS data sets.
- OUTLENGTH=n
- OUTL=n
specifies the length of those output variables that are not copied from the input data set but are created by PROC MODECLUS.
The OUTLENGTH= option applies only to the following variables that appear in all of the output data sets: _K_, _DK_, _CK_, _R_, _DR_, _CR_, _CASCAD_, _METHOD_, _NJOIN_, and _LOCAL_.
The minimum value is 2 or 3, depending on the operating system. The maximum value is 8. The default value is 8.
- POWER=n
- POW=n
specifies the power of the density used with the METHOD=6 option. The default value is 2.
- R=n
specifies the radius of the sphere of support for uniform-kernel density estimation and the neighborhood for clustering. Specifying R=n is equivalent to specifying both DR=n and CR=n. See the section Density Estimation for details.
- SHORT
- SIMPLE
- S
displays means, standard deviations, skewness, kurtosis, and a coefficient of bimodality. The SIMPLE option applies only to coordinate data.
- STANDARD
- STD
standardizes the variables to mean 0 and standard deviation 1. The STANDARD option applies only to coordinate data.
- SUM
uses sums for cascading density estimates. See the description of the CASCADE= option for details.
- TEST
performs approximate significance tests for the number of clusters. The R= or DR= option must also be specified with a nonzero value to obtain significance tests.
The significance tests performed by PROC MODECLUS are valid only for simple random samples, and they require at least 20 observations per cluster to have enough power to be of any use. See the section Significance Tests for details.
- THRESHOLD=n
- THR=n
specifies the assignment threshold used with the METHOD=6 option. The default is 0.5.
- TRACE
traces the process of cluster assignments when METHOD=6 is specified.
Copyright © SAS Institute, Inc. All Rights Reserved.