| The LOGISTIC Procedure |
Example 51.10 Overdispersion
In a seed germination test, seeds of two cultivars were planted in pots of two soil conditions. The following statements create the data set seeds, which contains the observed proportion of seeds that germinated for various combinations of cultivar and soil condition. The variable n represents the number of seeds planted in a pot, and the variable r represents the number germinated. The indicator variables cult and soil represent the cultivar and soil condition, respectively.
data seeds;
input pot n r cult soil;
datalines;
1 16 8 0 0
2 51 26 0 0
3 45 23 0 0
4 39 10 0 0
5 36 9 0 0
6 81 23 1 0
7 30 10 1 0
8 39 17 1 0
9 28 8 1 0
10 62 23 1 0
11 51 32 0 1
12 72 55 0 1
13 41 22 0 1
14 12 3 0 1
15 13 10 0 1
16 79 46 1 1
17 30 15 1 1
18 51 32 1 1
19 74 53 1 1
20 56 12 1 1
;
PROC LOGISTIC is used as follows to fit a logit model to the data, with cult, soil, and cult  soil interaction as explanatory variables. The option SCALE=NONE is specified to display goodness-of-fit statistics.
soil interaction as explanatory variables. The option SCALE=NONE is specified to display goodness-of-fit statistics.
proc logistic data=seeds;
model r/n=cult soil cult*soil/scale=none;
title 'Full Model With SCALE=NONE';
run;
Results of fitting the full factorial model are shown in Output 51.10.1. Both Pearson 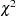 and deviance are highly significant (
and deviance are highly significant ( ), suggesting that the model does not fit well.
), suggesting that the model does not fit well.
| Model Fit Statistics | ||
|---|---|---|
| Criterion | Intercept Only |
Intercept and Covariates |
| AIC | 1256.852 | 1213.003 |
| SC | 1261.661 | 1232.240 |
| -2 Log L | 1254.852 | 1205.003 |
If the link function and the model specification are correct and if there are no outliers, then the lack of fit might be due to overdispersion. Without adjusting for the overdispersion, the standard errors are likely to be underestimated, causing the Wald tests to be too sensitive. In PROC LOGISTIC, there are three SCALE= options to accommodate overdispersion. With unequal sample sizes for the observations, SCALE=WILLIAMS is preferred. The Williams model estimates a scale parameter  by equating the value of Pearson
by equating the value of Pearson 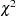 for the full model to its approximate expected value. The full model considered in the following statements is the model with cultivar, soil condition, and their interaction. Using a full model reduces the risk of contaminating
for the full model to its approximate expected value. The full model considered in the following statements is the model with cultivar, soil condition, and their interaction. Using a full model reduces the risk of contaminating  with lack of fit due to incorrect model specification.
with lack of fit due to incorrect model specification.
proc logistic data=seeds;
model r/n=cult soil cult*soil / scale=williams;
title 'Full Model With SCALE=WILLIAMS';
run;
Results of using Williams’ method are shown in Output 51.10.2. The estimate of  is 0.075941 and is given in the formula for the Weight Variable at the beginning of the displayed output.
is 0.075941 and is given in the formula for the Weight Variable at the beginning of the displayed output.
| Full Model With SCALE=WILLIAMS |
| Model Information | |
|---|---|
| Data Set | WORK.SEEDS |
| Response Variable (Events) | r |
| Response Variable (Trials) | n |
| Weight Variable | 1 / ( 1 + 0.075941 * (n - 1) ) |
| Model | binary logit |
| Optimization Technique | Fisher's scoring |
| Number of Observations Read | 20 |
|---|---|
| Number of Observations Used | 20 |
| Sum of Frequencies Read | 906 |
| Sum of Frequencies Used | 906 |
| Sum of Weights Read | 198.3216 |
| Sum of Weights Used | 198.3216 |
| Note: | Since the Williams method was used to accommodate overdispersion, the Pearson chi-squared statistic and the deviance can no longer be used to assess the goodness of fit of the model. |
| Model Fit Statistics | ||
|---|---|---|
| Criterion | Intercept Only |
Intercept and Covariates |
| AIC | 276.155 | 273.586 |
| SC | 280.964 | 292.822 |
| -2 Log L | 274.155 | 265.586 |
| Testing Global Null Hypothesis: BETA=0 | |||
|---|---|---|---|
| Test | Chi-Square | DF | Pr > ChiSq |
| Likelihood Ratio | 8.5687 | 3 | 0.0356 |
| Score | 8.4856 | 3 | 0.0370 |
| Wald | 8.3069 | 3 | 0.0401 |
| Analysis of Maximum Likelihood Estimates | |||||
|---|---|---|---|---|---|
| Parameter | DF | Estimate | Standard Error |
Wald Chi-Square |
Pr > ChiSq |
| Intercept | 1 | -0.3926 | 0.2932 | 1.7932 | 0.1805 |
| cult | 1 | -0.2618 | 0.4160 | 0.3963 | 0.5290 |
| soil | 1 | 0.8309 | 0.4223 | 3.8704 | 0.0491 |
| cult*soil | 1 | -0.0532 | 0.5835 | 0.0083 | 0.9274 |
Since neither cult nor cult  soil is statistically significant (
soil is statistically significant ( and
and  , respectively), a reduced model that contains only the soil condition factor is fitted, with the observations weighted by
, respectively), a reduced model that contains only the soil condition factor is fitted, with the observations weighted by  . This can be done conveniently in PROC LOGISTIC by including the scale estimate in the SCALE=WILLIAMS option as follows:
. This can be done conveniently in PROC LOGISTIC by including the scale estimate in the SCALE=WILLIAMS option as follows:
proc logistic data=seeds;
model r/n=soil / scale=williams(0.075941);
title 'Reduced Model With SCALE=WILLIAMS(0.075941)';
run;
Results of the reduced model fit are shown in Output 51.10.3. Soil condition remains a significant factor ( ) for the seed germination.
) for the seed germination.
Copyright © 2009 by SAS Institute Inc., Cary, NC, USA. All rights reserved.