| The HAPLOTYPE Procedure |
Example 8.4 Testing for Marker-Trait Associations
To demonstrate how the TRAIT statement can be used, a subset of data from GAW12 (Wijsman et al. 2001) is read into a SAS data set as follows:
data gaw;
input status $ a1-a24;
datalines;
U 8 4 4 4 2 7 3 2 1 4 10 2 6 6 1 2 1 1 7 7 8 7 8 8
U 5 9 3 5 3 4 2 3 4 3 14 10 3 6 7 7 1 4 5 12 3 3 1 2
A 8 2 5 1 6 3 3 5 3 4 5 3 3 1 5 3 3 4 7 7 7 3 7 7
U 7 8 5 3 8 4 5 3 3 4 13 8 1 3 4 5 4 4 10 7 1 2 2 2
U 9 2 2 5 7 6 9 3 2 4 3 2 5 2 1 2 2 4 5 7 4 3 1 12
U 2 7 1 4 6 7 8 4 4 3 10 5 5 2 4 3 3 1 8 11 2 3 7 7
U 7 7 6 6 1 4 9 5 3 1 14 6 5 3 1 3 3 1 12 1 3 7 7 7
U 4 4 3 7 3 2 8 9 3 1 9 10 6 4 5 3 1 4 10 8 8 5 8 2
A 8 9 6 5 6 4 3 4 4 1 9 1 7 7 2 5 4 1 1 1 5 1 10 2
U 9 5 6 1 2 6 3 3 3 2 8 7 1 5 3 8 1 3 1 8 3 5 1 4
U 8 1 1 5 8 6 3 3 4 3 1 10 3 1 2 3 4 4 5 10 4 5 7 9
A 7 2 3 4 1 3 2 3 3 3 7 1 7 7 2 3 3 4 5 1 5 5 7 9
U 9 3 1 1 2 3 9 8 3 1 13 13 7 1 2 2 3 4 10 3 1 1 10 1
U 2 9 6 1 3 4 3 2 4 3 2 1 4 3 8 1 4 3 9 5 4 2 1 10
U 2 1 1 4 4 7 5 8 3 4 10 13 5 4 4 4 4 3 12 2 3 7 2 12
U 7 7 6 6 3 3 9 3 4 3 14 14 2 1 2 2 1 4 9 1 5 8 4 10
U 1 3 6 5 5 4 9 4 3 4 13 1 2 3 1 2 1 3 1 3 5 3 2 1
U 9 2 6 6 3 4 3 4 2 4 14 9 5 2 4 4 1 1 12 7 5 5 11 7
U 3 3 5 5 8 4 6 5 4 3 2 13 7 1 1 2 3 2 10 7 3 4 7 10
U 4 3 4 5 7 7 8 8 3 3 8 13 3 4 3 2 4 1 1 12 1 3 10 7
U 3 8 1 1 3 8 8 3 4 4 13 12 1 4 5 7 1 4 1 8 3 2 3 3
U 7 8 5 7 7 3 3 3 4 3 14 5 5 1 8 5 4 4 12 12 5 5 10 10
A 7 2 5 4 1 3 3 9 4 3 13 9 2 3 6 5 4 4 1 10 5 2 1 10
U 7 2 4 5 6 1 1 2 4 4 10 8 4 5 5 4 1 1 6 9 2 7 2 12
U 3 3 4 2 7 3 8 3 4 4 14 12 3 2 5 4 3 3 9 3 2 1 12 12
A 2 3 4 1 4 3 3 3 4 4 6 14 1 1 2 2 1 3 3 1 2 8 2 7
U 5 9 3 1 7 4 3 4 2 4 9 8 5 7 3 1 1 3 9 9 2 5 1 9
U 8 5 6 5 3 7 4 4 4 3 10 9 7 5 2 8 4 1 7 8 2 7 12 1
U 9 8 5 5 7 3 6 5 1 3 13 5 2 2 8 7 3 3 9 12 1 3 4 1
A 7 8 5 2 3 5 3 9 3 3 12 5 1 1 1 2 1 4 7 2 5 3 6 1
A 5 4 1 1 3 7 4 5 3 3 14 13 7 3 3 1 4 3 1 8 3 3 2 9
U 8 9 3 2 7 3 8 9 4 1 1 12 5 4 4 6 3 4 2 7 5 2 3 10
A 9 2 3 5 3 3 2 3 2 3 14 13 6 1 3 1 4 3 3 2 3 1 1 7
A 2 5 7 5 6 7 9 4 3 4 14 13 5 1 2 3 4 4 2 10 3 1 12 12
U 7 2 3 1 1 3 4 4 3 4 2 8 5 3 4 6 3 3 10 12 8 3 2 1
A 7 5 1 5 3 3 9 2 3 3 10 6 1 7 2 4 4 4 10 9 1 8 7 3
U 3 2 5 5 4 3 3 5 1 3 1 1 5 2 1 2 3 3 10 3 3 3 10 4
A 3 2 5 5 8 5 3 7 4 3 2 14 5 5 3 3 3 4 11 1 6 2 1 10
A 2 7 5 5 3 2 9 4 3 3 1 7 7 5 4 7 4 1 12 7 2 3 12 9
A 5 7 2 3 7 3 3 3 3 4 9 2 4 1 2 7 1 4 6 1 2 1 7 7
U 7 4 3 4 5 3 3 8 3 3 2 8 4 6 7 7 4 1 3 1 2 4 12 1
U 7 8 5 4 4 7 9 9 4 3 5 13 7 1 4 4 4 4 9 8 8 3 3 10
U 2 8 4 5 3 7 3 4 3 3 8 14 6 4 6 2 3 4 7 1 3 3 3 10
U 6 8 1 3 6 7 5 4 3 4 1 12 3 7 8 4 3 4 12 12 4 7 12 6
A 8 7 3 1 3 6 4 4 3 3 4 10 6 5 8 1 1 4 1 10 2 2 5 2
U 2 8 6 6 4 8 4 3 4 3 9 1 1 1 2 3 4 4 2 6 2 3 9 7
U 9 8 4 3 7 3 8 4 4 3 8 8 6 6 4 5 3 4 5 5 1 8 10 1
U 9 3 5 1 8 6 5 3 3 2 13 2 3 5 8 2 1 3 1 10 3 3 10 12
U 2 9 1 6 7 4 9 9 4 1 8 1 3 2 5 8 4 4 3 1 3 3 12 7
U 8 8 6 2 3 2 2 4 3 4 6 12 3 1 7 2 4 4 5 9 2 3 1 10
;
This data set contains 12 markers. Suppose you are interested in testing three of the marker loci at a time for association with the trait (status in this case: "A" for affected or "U" for unaffected with a particular disease) over all of their haplotypes. That is, assuming the markers are numbered in the order in which they appear on the chromosome, haplotypes at marker loci 1 through 3 are analyzed, then haplotypes at marker loci 4 through 6 are analyzed, and so on. These tests can be performed in addition to, or in place of, single-marker case-control tests (see Chapter 5 for more information). In order to reduce the amount of SAS code needed for this analysis, a SAS macro can be used as follows:
%macro hap_trait;
%do firsta=1 %to 19 %by 6;
%let lasta=%eval(&firsta+5);
%let firstm=%eval((&firsta+1)/2);
%let lastm=%eval(&lasta/2);
proc haplotype data=gaw noprint;
var a&firsta-a&lasta;
trait status;
run;
%end;
%mend;
%hap_trait
Since the NOPRINT option is specified, this code produces only the "Test for Marker-Trait Association" table each of the four times PROC HAPLOTYPE is invoked.
| Test for Marker-Trait Association | ||||||
|---|---|---|---|---|---|---|
| Trait Number |
Trait Value |
Num Obs |
DF | LogLike | Chi-Square | Pr > ChiSq |
| 1 | U | 36 | 156 | -245.18487 | ||
| 2 | A | 14 | 68 | -69.90500 | ||
| Combined | 50 | 181 | -355.16139 | 80.1430 | 0.0005 | |
| Test for Marker-Trait Association | ||||||
|---|---|---|---|---|---|---|
| Trait Number |
Trait Value |
Num Obs |
DF | LogLike | Chi-Square | Pr > ChiSq |
| 1 | U | 36 | 140 | -236.78471 | ||
| 2 | A | 14 | 62 | -78.22280 | ||
| Combined | 50 | 162 | -349.30084 | 68.5867 | 0.0033 | |
Output 8.4.1 displays the four tables that are created by this macro. The first corresponds to testing the three-locus haplotypes at the first three marker loci with the TRAIT variable, the second corresponds to the second set of three markers, and so on. From the LRTs that are performed and summarized in the output, it can be concluded that out of the four sets of marker loci tested, the haplotypes at markers 10, 11, and 12 show the most significant association with the trait variable status. The chi-square statistic for testing the haplotypes at these markers for association with disease status is calculated as 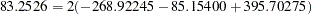 with degrees of freedom
with degrees of freedom 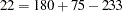 , which has a
, which has a  -value < 0.0001.
-value < 0.0001.
Suppose you want to further explore the association between these three markers and the trait. You can also perform tests of association between each individual haplotype at these marker loci and disease status by using the following code:
ods output haplotype.haptraittest=outhap;
proc haplotype data=gaw noprint;
var a19-a24;
trait status / testall perms=100;
run;
proc print data=outhap(obs=20) noobs;
title 'The HAPLOTYPE Procedure';
title2 ' ';
title3 'Tests for Haplotype-Trait Association';
run;
The TESTALL option indicates that a test for trait association should be performed on each haplotype by using a chi-square test statistic, which is performed by default. In addition, since the PERMS=100 option is included, an empirical  -value is calculated. Because of the number of alleles at each marker in this example, this option increases the computation time substantially, even with this small number of permutations.
-value is calculated. Because of the number of alleles at each marker in this example, this option increases the computation time substantially, even with this small number of permutations.
| The HAPLOTYPE Procedure |
| Tests for Haplotype-Trait Association |
| Number | Haplotype | Trait1Freq | Trait2Freq | CombinedFreq | ChiSq | ProbChiSq | ProbExact |
|---|---|---|---|---|---|---|---|
| 1 | 1-1-2 | 0.00000 | 0.03571 | 0.00000 | 0 | 1.0000 | 1.0000 |
| 2 | 1-1-7 | 0.00000 | 0.00000 | 0.01000 | 1.0101 | 0.3149 | 0.3800 |
| 3 | 1-1-10 | 0.00000 | 0.00000 | 0.01950 | 1.9883 | 0.1585 | 0.3400 |
| 4 | 1-2-1 | 0.00000 | 0.01786 | 0.03000 | 2.3686 | 0.1238 | 0.3000 |
| 5 | 1-2-2 | 0.00000 | 0.05357 | 0.01000 | 6.0967 | 0.0135 | 0.0500 |
| 6 | 1-2-3 | 0.00000 | 0.00000 | 0.00000 | 0.001666 | 0.9674 | 0.6500 |
| 7 | 1-2-5 | 0.00000 | 0.00000 | 0.01000 | 1.0101 | 0.3149 | 0.3400 |
| 8 | 1-2-7 | 0.00000 | 0.05357 | 0.00000 | 0 | 1.0000 | 1.0000 |
| 9 | 1-2-10 | 0.00000 | 0.01786 | 0.00000 | 0 | 1.0000 | 1.0000 |
| 10 | 1-2-12 | 0.00000 | 0.00000 | 0.00000 | 0 | 1.0000 | 1.0000 |
| 11 | 1-3-1 | 0.00694 | 0.00000 | 0.00000 | 0 | 1.0000 | 1.0000 |
| 12 | 1-3-2 | 0.00000 | 0.01786 | 0.01000 | 0.9019 | 0.3423 | 0.4000 |
| 13 | 1-3-3 | 0.02777 | 0.00000 | 0.02000 | 0.7934 | 0.3731 | 0.7700 |
| 14 | 1-3-4 | 0.00000 | 0.00000 | 0.00000 | 0 | 1.0000 | 1.0000 |
| 15 | 1-3-7 | 0.04167 | 0.00000 | 0.02045 | 2.2035 | 0.1377 | 0.1000 |
| 16 | 1-3-9 | 0.00000 | 0.01786 | 0.00000 | 0 | 1.0000 | 1.0000 |
| 17 | 1-3-10 | 0.00000 | 0.00000 | 0.00000 | 7.8011E-8 | 0.9998 | 0.9500 |
| 18 | 1-3-12 | 0.01389 | 0.00000 | 0.01006 | 0.3905 | 0.5320 | 0.8700 |
| 19 | 1-4-1 | 0.01389 | 0.00000 | 0.00000 | 0 | 1.0000 | 1.0000 |
| 20 | 1-4-12 | 0.00000 | 0.00000 | 0.00000 | 0 | 1.0000 | 1.0000 |
Output 8.4.2 displays the table "Test for Haplotype-Trait Association" as a SAS data set by using the ODS system in order to show only the first 20 rows. The table contains haplotypes at markers 10, 11, and 12 and their estimated frequencies among individuals with the first trait value, individuals with the second trait value, and all individuals. The chi-square statistic testing whether the frequencies between the two trait groups are significantly different is also shown, along with its 1 df  -value. Note that none of the haplotypes shown here have an association with disease status significant at the 0.05 level according to the approximations of exact
-value. Note that none of the haplotypes shown here have an association with disease status significant at the 0.05 level according to the approximations of exact  -values.
-values.
Copyright © 2008 by SAS Institute Inc., Cary, NC, USA. All rights reserved.