Sample 45566: Bayesian Exponential Mixture Model
 |  |  |  |
PDF of Example | SAS/STAT Focus Area Examples
Analysis
The LOS data analyzed in this example originate from geriatric patients in a psychiatric hospital in North East London in 1991 and were studied by Harrison and Millard ( 1991 ) and McClean and Millard ( 1993 ) . Each observation represents the LOS in days for an admitted patient.
data inputdata;
input los @@;
datalines;
1671 1300 722 586 552 525 364
359 321 302 272 248 226 216
208 182 141 141 132 120 117
115 114 113 104 103 101 99
96 94 93 92 88 84 83
81 79 74 70 63 62 62
61 56 55 53 53 51 51
50 49 36 33 33 33 29
28 26 24 19 16 16 15
... more lines ...
35 28 20 19 9 5 2
2 2 22317 14006 11549 11006 8981
8402 7947 7266 6693 4408 4010 4003
1970 1857 1849 1833 1770 1769 1514
1217 956 924 611 386 280 93
;
Before using this data in a mixture model setting, use the MEANS procedure to obtain summary statistics for the LOS data in table form. Figure 1 displays summary statistics for this data.
| Analysis Variable : los | ||||
|---|---|---|---|---|
| N | Mean | Std Dev | Minimum | Maximum |
| 469 | 3712.36 | 5675.97 | 1.0000000 | 24028.00 |
Figure 2 displays a plot that illustrates the skewness of the data. The histogram of the data is overlaid with a scaled exponential density, and you can see that a single exponential density does not fit the lower values of LOS well.

Bayesian Mixture of Exponential Model
Mixture models arise naturally when one mechanism generates data according to one model and another mechanism generates data according to a different model. In this data set, the variable that indicates which mechanism generated an observation is not recorded, and only the response variable is available. A mixture model is useful for analyzing this data set because it unveils the latent heterogeneity that arises from a latent categorical variable (Fruhwirth-Schnatter; 2006 ) .
Mixture models can be expressed as a weighted average of  component densities. More formally,
component densities. More formally,  is said to arise from a finite mixture model with probability density function
is said to arise from a finite mixture model with probability density function  where
where
 |
For all  ,
,  is the component probability density function, and
is the component probability density function, and  are the weights defined by the following constraints:
are the weights defined by the following constraints:  and
and  .
.
Congdon ( 2003 ) states that exponential mixture models with relatively small number of components are effective in modeling skewed LOS data. You can write a two-component exponential mixture model for LOS data with density as follows:
 |
for patients  .
.
There are three parameters in the density:  , and
, and  . Researchers Harrison and Millard ( 1991 ) suggest the mean parameters,
. Researchers Harrison and Millard ( 1991 ) suggest the mean parameters,  and
and  , as the average LOS for the standard- and long-stay groups, respectively. In addition,
, as the average LOS for the standard- and long-stay groups, respectively. In addition,  represents an unknown fraction of patients in the standard-stay group with
represents an unknown fraction of patients in the standard-stay group with  .
.
Suppose the following prior distributions are placed on the three parameters:
 |
 |
 |
|||
 |
 |
 |
where  indicates a prior distribution and
indicates a prior distribution and  is the density function for the inverse-gamma distribution. Priors of this type are often called diffuse priors . The
is the density function for the inverse-gamma distribution. Priors of this type are often called diffuse priors . The  prior expresses your lack of knowledge about the mixture proportion.
prior expresses your lack of knowledge about the mixture proportion.
Label-switching is a common problem that arises in mixture models. It was described by as a result of the invariance of the mixture likelihood function under the relabeling of the mixture components. In an effort to remove label-switching, Harrison and Millard ( 1991 ) place an identifiability constraint that the average LOS of the standard-stay group is less than that of the long-stay group; that is,  .
.
Using Bayes’ theorem, the likelihood function and prior distributions determine the posterior distribution of  , and
, and  as follows:
as follows:
 |
PROC MCMC obtains samples from the desired posterior distribution, which is determined by the prior and likelihood specified. It does not require the form of the posterior distribution.
The following SAS statements use the prior distributions to fit the Bayesian exponential mixture model. The PROC MCMC statement invokes the procedure and specifies the input data set. The NMC= option specifies the number of posterior simulation iterations. The THIN=5 option specifies that one of every five samples is kept. The PROPCOV=QUANEW option uses the estimated inverse Hessian matrix as the initial proposal covariance matrix.
ods graphics on;
proc mcmc data=inputdata seed=1010 nmc=50000 thin=5
propcov=quanew statistics=summary;
ods output PostSummaries = post_summ;
parms B 100 D 6000 pi 0.5;
prior B D ~ igamma(3/10, scale = 10/3);
prior pi ~ uniform(0,1);
if (B < D) then
llike = log(pi*pdf("expo", los, B) + (1-pi)*pdf("expo", los, D));
else
llike = .;
model general(llike);
run;
ods graphics off;
The ODS OUTPUT statement creates an output data set post_summ used later for a graphical analysis of the fit in Figure 8 . The PARMS statement puts all three parameters  ,
,  , and
, and  in a single block and assigns initial values to each of them. The PRIOR statements specify priors for all the parameters. Note that
in a single block and assigns initial values to each of them. The PRIOR statements specify priors for all the parameters. Note that  and
and  can be specified with one PRIOR statement because they have the same prior distribution.
can be specified with one PRIOR statement because they have the same prior distribution.
The IF-ELSE statements enable different values of LOS to have different log-likelihood functions, depending on whether the order constraint placed on the mean parameters is satisfied. The MODEL statement specifies that llike is the log likelihood for each observation in the model and is simply a missing value when the order constraint is not met.
By turning ODS Graphics on, PROC MCMC produces graphs at the end of the procedure which enable you to visually examine the convergence of the chain. See Figure 3 . Inferences should not be made if the Markov chain has not converged.
 ,
,  , and
, and 
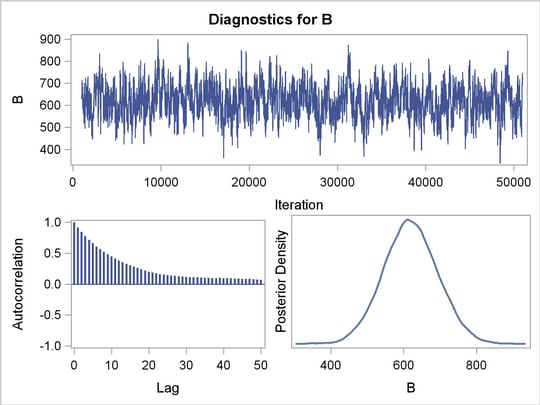


Figure 3 displays convergence diagnostic graphs for parameters  ,
,  , and
, and  . The trace plots indicate that the chains appear to have reached stationary distributions. The chain also has good mixing and is dense.
. The trace plots indicate that the chains appear to have reached stationary distributions. The chain also has good mixing and is dense.
The autocorrelation plots indicate low autocorrelation and efficient sampling. Finally, the kernel density plots show the smooth, unimodal shape of posterior marginal distributions for each parameter. If label-switching had occured, you would see jumps in the trace plots or multimodal density plots.
Figure 4 contains the "Parameters" table which lists the names of the parameters, the blocking information, the sampling method used, the starting values, and the prior distributions.
| Parameters | |||
|---|---|---|---|
| Parameter | Sampling Method |
Initial Value |
Prior Distribution |
| B | N-Metropolis | 100.0 | igamma(3/10, scale = 10/3) |
| D | N-Metropolis | 6000.0 | igamma(3/10, scale = 10/3) |
| pi | N-Metropolis | 0.5000 | uniform(0,1) |
The "Tuning History" table, shown in Figure 5 , displays how the tuning stage progresses for the multivariate random walk Metropolis algorithm used by PROC MCMC to generate samples from the posterior distribution. An important aspect of the algorithm is the calibration of the proposal distribution. The tuning of the Markov chain is broken into a number of phases. In each phase, PROC MCMC generates trial samples and automatically modifies the proposal distribution as a result of the acceptance rate.
The "Burn-In History" and the "Sampling History" tables show the burn-in and main phase sampling, respectively.
| Tuning History | |||
|---|---|---|---|
| Phase | Block | Scale | Acceptance Rate |
| 1 | 1 | 2.3800 | 0.0280 |
| 2 | 1 | 1.0123 | 0.0700 |
| 3 | 1 | 0.5221 | 0.3420 |
| Burn-In History | ||
|---|---|---|
| Block | Scale | Acceptance Rate |
| 1 | 0.5221 | 0.3980 |
| Sampling History | ||
|---|---|---|
| Block | Scale | Acceptance Rate |
| 1 | 0.5221 | 0.3811 |
Figure 6 displays summary and interval statistics for each posterior distribution.
| Posterior Summaries | ||||||
|---|---|---|---|---|---|---|
| Parameter | N | Mean | Standard Deviation |
Percentiles | ||
| 25% | 50% | 75% | ||||
| B | 10000 | 619.1 | 74.2261 | 568.9 | 618.3 | 668.3 |
| D | 10000 | 7752.6 | 721.2 | 7267.1 | 7710.4 | 8202.1 |
| pi | 10000 | 0.5638 | 0.0403 | 0.5370 | 0.5655 | 0.5916 |
| Posterior Intervals | |||||
|---|---|---|---|---|---|
| Parameter | Alpha | Equal-Tail Interval | HPD Interval | ||
| B | 0.050 | 473.6 | 767.0 | 472.0 | 763.7 |
| D | 0.050 | 6418.1 | 9288.4 | 6339.5 | 9195.0 |
| pi | 0.050 | 0.4826 | 0.6389 | 0.4825 | 0.6387 |
Figure 7 reports a number of convergence diagnostics to assist in determining convergence. These are the Monte Carlo standard errors, the autocorrelations at selected lags, the Geweke diagnostics, and the effective sample sizes.
| Monte Carlo Standard Errors | |||
|---|---|---|---|
| Parameter | MCSE | Standard Deviation |
MCSE/SD |
| B | 3.9598 | 74.2261 | 0.0533 |
| D | 38.2006 | 721.2 | 0.0530 |
| pi | 0.00164 | 0.0403 | 0.0408 |
| Posterior Autocorrelations | ||||
|---|---|---|---|---|
| Parameter | Lag 1 | Lag 5 | Lag 10 | Lag 50 |
| B | 0.9150 | 0.6588 | 0.4459 | 0.0689 |
| D | 0.9220 | 0.6766 | 0.4760 | 0.0354 |
| pi | 0.5086 | 0.3438 | 0.2564 | 0.0267 |
| Geweke Diagnostics | ||
|---|---|---|
| Parameter | z | Pr > |z| |
| B | 0.1718 | 0.8636 |
| D | -0.2669 | 0.7895 |
| pi | -0.3073 | 0.7586 |
| Effective Sample Sizes | |||
|---|---|---|---|
| Parameter | ESS | Correlation Time |
Efficiency |
| B | 351.4 | 28.4595 | 0.0351 |
| D | 356.4 | 28.0580 | 0.0356 |
| pi | 601.9 | 16.6129 | 0.0602 |
Figure 6 displays the marginal posterior summaries. The larger group of standard-stay patients has an average LOS of  days relative to the smaller group of long-stay patients whose stay averages
days relative to the smaller group of long-stay patients whose stay averages  days. The posterior average mixture proportions are
days. The posterior average mixture proportions are  and
and  for the standard-stay and long-stay group, respectively.
for the standard-stay and long-stay group, respectively.
Figure 8 illustrates the scaled mixture of exponential density and the gain in model fit from the two-component exponential mixture model.

The single-component exponential model had an average LOS of 3712 days. The mean parameters found when fitting an exponential mixture model to the standard-stay group and the long-stay group are 619 and 7752 days, respectively. The mixture distribution fits the data better than the exponential distribution, especially at the low values of LOS. Additional components could be fit and evaluated in similar methods or with information criteria.
References
-
Congdon, P. (2003), Applied Bayesian Modeling , John Wiley & Sons.
-
Fruhwirth-Schnatter, S. (2006), Finite Mixture and Markov Switching Models , Springer.
-
Harrison, G. and Millard, P. (1991), “Balancing Acute and Long-Term Care: The Mathematics of Throughput in Departments of Geriatric Medicine,” Meth. Infor. Medicine , 30, 221–228.
-
McClean, S. and Millard, P. (1993), “Patterns of Length of Stay after Admission in Geriatric Medicine: An Event History Approach,” The Statistician , 42(3), 263–274.
These sample files and code examples are provided by SAS Institute Inc. "as is" without warranty of any kind, either express or implied, including but not limited to the implied warranties of merchantability and fitness for a particular purpose. Recipients acknowledge and agree that SAS Institute shall not be liable for any damages whatsoever arising out of their use of this material. In addition, SAS Institute will provide no support for the materials contained herein.
data inputdata;
input los @@;
datalines;
1671 1300 722 586 552 525 364
359 321 302 272 248 226 216
208 182 141 141 132 120 117
115 114 113 104 103 101 99
96 94 93 92 88 84 83
81 79 74 70 63 62 62
61 56 55 53 53 51 51
50 49 36 33 33 33 29
28 26 24 19 16 16 15
14 9 9 5 5 3 3
2 2 1 24028 23871 22852 22464
22317 21525 21264 21131 21077 20944 20608
20133 19583 19038 18821 18761 18742 18479
18449 18241 18128 17830 17800 17746 17285
16763 16444 15922 15226 15224 15178 14885
14744 14715 14601 14482 14174 14149 13295
13274 12850 12661 12660 12405 12300 12248
11821 11769 11577 11159 10680 10414 10393
10253 10095 9926 9722 9670 9426 9389
9109 9004 9003 8959 8927 8583 8465
8464 8317 8176 8156 8114 8096 7884
7639 6552 6387 6283 6243 6155 5917
5397 5370 5165 5007 4899 4408 4395
4290 4282 4208 4182 4153 3855 3810
3735 3719 3676 3467 3225 3149 3115
2989 2933 2927 2918 2802 2746 2701
2542 2430 2421 2380 2340 2331 2318
2235 2209 2053 1997 1978 1923 1856
1812 1804 1792 1757 1717 1670 1666
1661 1615 1610 1609 1525 1481 1424
1388 1376 1349 1302 1159 1134 1057
1006 937 756 733 723 588 533
522 348 212 439 188 181 141
131 116 96 83 56 40 40
37 36 26 23 19 14 8
20809 20319 18918 15058 13540 13446 10161
9163 6474 5420 4584 4341 3805 3640
3640 3578 3453 3375 3186 2756 2724
2658 2599 2583 2416 2261 2232 2127
2126 2035 2015 1986 1903 1897 1874
1780 1776 1759 1752 1749 1730 1624
1580 1552 1549 1546 1493 1472 1469
1456 1441 1433 1428 1399 1391 1378
1363 1359 1317 1295 1279 1262 1247
1201 1199 1195 1182 1175 1141 1135
1106 1049 1037 1031 1027 1006 994
972 952 944 938 938 936 933
897 887 880 880 863 842 806
792 791 789 764 762 726 712
707 706 703 697 691 687 677
649 635 613 607 602 602 601
594 590 576 569 552 544 544
539 532 530 525 511 509 502
497 477 462 453 446 443 434
426 414 411 407 399 397 391
375 374 370 366 365 359 357
351 350 348 345 343 341 337
336 336 333 330 330 310 306
300 289 285 273 260 252 238
231 223 219 215 197 197 197
187 187 161 126 125 123 107
100 93 83 58 48 37 37
35 28 20 19 9 5 2
2 2 22317 14006 11549 11006 8981
8402 7947 7266 6693 4408 4010 4003
1970 1857 1849 1833 1770 1769 1514
1217 956 924 611 386 280 93
;
/*-----------------------------------------------------------------
Example: Fitting a Mixture of Exponential Distributions for
Patients Length of Stay
Requires: SAS/STAT
Version: 9.2
------------------------------------------------------------------*/
** Calculates the Average number of days in hospital stays **;
proc means data = inputdata;
ods output Summary=summ;
run;
** Creates a macro variable for the mean LOS **;
data _null_;
set summ;
call symputx("los_Mean",los_Mean);
run;
** Calculates the Scaled density of Exponential **;
data a;
do x = 10 to 30000 by 1;
ex1 = 300*pdf("expo", x, &los_Mean);
output;
end;
run;
** Merges inputdata and scaled density data set **;
data a_inputdata;
set a inputdata;
run;
** Template for displaying Histogram, and both scaled density plots **;
proc template;
define statgraph sasuser.hist;
begingraph;
entrytitle "Distribution and Scaled Posterior Densities of Length of Stays";
layout overlay / border=FALSE
xaxisopts=(label="Length of Stay (days)")
yaxisopts=(label="Scaled Density");
seriesplot x = x y = ex1 / primary = true
lineattrs=(thickness = 2 color=red)
name = '1' legendlabel = "Exponential Density";
histogram los / binwidth=300 scale=proportion
fillattrs=(color=lightgray transparency = .6);
discretelegend '1'/across=2 autoalign=(center)
location=outside;
endlayout;
endgraph;
end;
run;
ods graphics on;
** Create graph **;
proc sgrender data=a_inputdata template=sasuser.hist;
run;
ods graphics off;
ods graphics on;
proc mcmc data=inputdata seed=1010 nmc=50000 thin=5
propcov=quanew;
ods output PostSummaries = post_summ;
parms B 100 D 6000 pi 0.5;
prior B D ~ igamma(3/10, scale = 10/3);
prior pi ~ uniform(0,1);
if (B < D) then
llike = log(pi*pdf("expo", los, B) + (1-pi)*pdf("expo", los, D));
else
llike = .;
model general(llike);
run;
ods graphics off;
** Creates several macro variables for calculating posterior density **;
data _null_;
set post_summ;
if(Parameter = "B") then do;
b_temp = Mean;
call symputx("Bmean",b_temp);
end;
if(Parameter = "D") then do;
d_temp = Mean;
call symputx("Dmean",d_temp);
end;
if(Parameter = "pi") then do;
pi_temp = Mean;
call symputx("pimean",pi_temp);
end;
run;
** Creates ex2 variable, scaled density values of mixture **;
data b;
do x2 = 200 to 30000 by 1;
ex2 = 300*(&pimean*pdf("expo", x2, &Bmean) +
(1- &pimean)*pdf("expo", x2, &Dmean));
output;
end;
run;
** Creates combined output data sets **;
data b_inputdata;
set a b inputdata;
run;
** Template for displaying Histogram, and scaled density plots **;
proc template;
define statgraph sasuser.hist2;
begingraph;
entrytitle "Distribution and Scaled Posterior Densities of Length of Stays";
layout overlay / border=FALSE
xaxisopts=(label="Length of Stay (days)")
yaxisopts=(label="Scaled Density");
seriesplot x = x y = ex1 /primary = true
lineattrs=(thickness = 2 color=red)
name = '1' legendlabel = "Exponential Density";
seriesplot x = x2 y = ex2 /primary = true
lineattrs=(thickness = 2 color=blue)
name = '2' legendlabel = "Mixture Density";
histogram los/binwidth=300 scale=proportion
fillattrs=(color=lightgray transparency = .6);
discretelegend '1' '2'/across=2 autoalign=(center)
location=outside;
endlayout;
endgraph;
end;
run;
ods graphics on;
** Create graph **;
proc sgrender data=b_inputdata template=sasuser.hist2;
run;
ods graphics off;
These sample files and code examples are provided by SAS Institute Inc. "as is" without warranty of any kind, either express or implied, including but not limited to the implied warranties of merchantability and fitness for a particular purpose. Recipients acknowledge and agree that SAS Institute shall not be liable for any damages whatsoever arising out of their use of this material. In addition, SAS Institute will provide no support for the materials contained herein.
| Type: | Sample |
| Topic: | Analytics ==> Bayesian Analysis SAS Reference ==> Procedures ==> MCMC |
| Date Modified: | 2018-05-31 10:19:33 |
| Date Created: | 2012-02-02 15:32:25 |
Operating System and Release Information
| Product Family | Product | Host | Product Release | SAS Release | ||
| Starting | Ending | Starting | Ending | |||
| SAS System | SAS/STAT | z/OS | 9.3 | |||
| Microsoft® Windows® for x64 | 9.3 | |||||
| Microsoft Windows 95/98 | 9.3 | |||||
| Microsoft Windows 2000 Advanced Server | 9.3 | |||||
| Microsoft Windows 2000 Datacenter Server | 9.3 | |||||
| Microsoft Windows 2000 Server | 9.3 | |||||
| Microsoft Windows 2000 Professional | 9.3 | |||||
| Microsoft Windows NT Workstation | 9.3 | |||||
| Microsoft Windows Server 2003 Datacenter Edition | 9.3 | |||||
| Microsoft Windows Server 2003 Enterprise Edition | 9.3 | |||||
| Microsoft Windows Server 2003 Standard Edition | 9.3 | |||||
| Microsoft Windows Server 2003 for x64 | 9.3 | |||||
| Microsoft Windows Server 2008 | 9.3 | |||||
| Microsoft Windows Server 2008 for x64 | 9.3 | |||||
| Microsoft Windows XP Professional | 9.3 | |||||
| Windows 7 Enterprise 32 bit | 9.3 | |||||
| Windows 7 Enterprise x64 | 9.3 | |||||
| Windows 7 Home Premium 32 bit | 9.3 | |||||
| Windows 7 Home Premium x64 | 9.3 | |||||
| Windows 7 Professional 32 bit | 9.3 | |||||
| Windows 7 Professional x64 | 9.3 | |||||
| Windows 7 Ultimate 32 bit | 9.3 | |||||
| Windows 7 Ultimate x64 | 9.3 | |||||
| Windows Millennium Edition (Me) | 9.3 | |||||
| Windows Vista | 9.3 | |||||
| Windows Vista for x64 | 9.3 | |||||
| 64-bit Enabled AIX | 9.3 | |||||
| 64-bit Enabled HP-UX | 9.3 | |||||
| 64-bit Enabled Solaris | 9.3 | |||||
| HP-UX IPF | 9.3 | |||||
| Linux | 9.3 | |||||
| Linux for x64 | 9.3 | |||||
| Solaris for x64 | 9.3 | |||||