Usage Note 37228: Estimating the difference in event probability (risk difference or marginal effect) with confidence interval
 |  |  |
Since the log odds (also called the logit) is the response function in a logistic model, such models enable you to estimate the log odds for populations in the data. A population is a setting of the model predictors. By exponentiating you can estimate the odds. Similarly, the difference between two populations results in an estimated difference in log odds that is equivalent to a log odds ratio. Again, by exponentiating you can estimate the odds ratio comparing the populations. So, simple linear combinations of the model parameters, such as can be specified in or generated by PROC LOGISTIC using the ODDSRATIO, LSMEANS, LSMESTIMATE, SLICE, CONTRAST, or ESTIMATE statements, can provide estimates of odds and odds ratios.
However, the risk difference or difference in population event probabilities (population means) cannot be obtained in this way. To estimate the difference in probabilities, you need to estimate a nonlinear function of the parameters of the logistic model. Alternatively, you could model the probabilities themselves rather than the log odds.
Several methods are illustrated in this note showing how the difference in population probabilities can be estimated and tested in logistic models with either a binary, or a multinomial response. The simplest method uses the Margins macro (SAS Note 63038) to fit the model, estimate the individual population probabilities (the predictive margins), and the difference in probabilities (the marginal effect).
When the variable whose risk difference is to be estimated is involved in interaction in a model, the risk difference changes depending on the level of the interacting variable(s). When the interacting variable is continuous, the risk difference can be estimated at various levels of the interacting variable and plotted as a function of the interacting variable as discussed in SAS Note 69621.
Other notes illustrate related tasks. To estimate the difference in probabilities with matched pairs data, see SAS Note 46997. For estimating the odds ratio with matched pairs data, see SAS Note 23127. For the simple case of a 2×2 table, a nonmodeling approach to estimating the difference in probabilities can be taken using PROC FREQ as described in SAS Note 22561.
The following data are used to illustrate the various methods of estimating and testing the difference in probabilities. Data set TEST has 100 observations containing two variables: a 0,1 coded binary response variable, Y, with 1 representing the event of interest, and a categorical predictor variable, A, with levels 1, 2, and 3. The following table summarizes the data.
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||
Binary Response — Using the Margins macro
See the description of the Margins macro for information on making it available in your SAS® session.
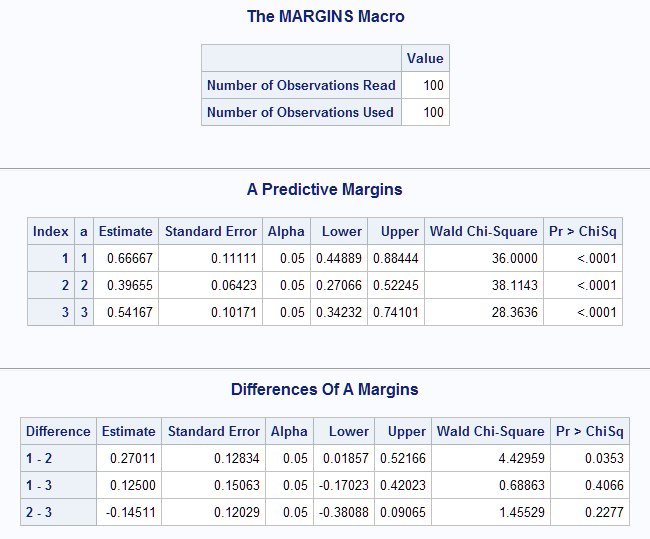
The following call of the Margins macro fits a logistic model and then estimates the predictive margins (probability that Y=1 at each level of A). A test that each equals zero is provided. Differences among the probabilities (marginal effects) are requested with the diff option. The cl option requests confidence intervals for the margins and the differences.
%Margins(data = test,
response = y,
roptions = event='1',
class = a,
model = a,
dist = binomial,
margins = a,
options = diff cl)
The estimated event probabilities at the levels of A are shown in the Predictive Margins table. The pairwise differences are given in the following table. Note that only the difference between the A=1 and A=2 levels (0.27) is significant (p=0.0353).
 |
Binary Response — Using the NLMeans macro
See the description of the NLMeans macro (SAS Note 62362) for information on making it available in your SAS® session.
The following statements fit a logistic model and the probabilities for the three A populations are estimated by the ILINK option in the LSMEANS statement. Adding the DIFF option in the LSMEANS statement would estimate the differences in log odds (equivalently, the log odds ratios), not the differences in the probabilities. The E option displays the table of coefficients that defines the LS-means for each level of A. The STORE statement saves the fitted model in an item-store named LogFit. The ODS OUTPUT statement saves the LSMEANS coefficients table in a data set named Coeffs.
proc logistic data=test;
class a / param=glm;
model y(event="1") = a;
lsmeans a / e ilink;
store LogFit;
ods output coef=Coeffs;
run;
In the table displayed by the LSMEANS statement, the estimated population probabilities appear in the column labeled Mean. Notice that the difference in the predicted probabilities for levels 1 and 2 of A is approximately 0.6666 - 0.3966 = 0.2700.
|
|||||||||||||||||||||||||||||||||||
The following call of the NLMeans macro estimates all pairwise differences among the probabilities of the three A populations. Specify the saved model with instore= and the saved LSMEANS coefficients with coef=. Since the link function in a logistic model is the logit, that is specified in link=. An optional title is specified in title=.
%NLMeans(instore=LogFit, coef=Coeffs, link=logit,
title=Differences of A Means)
The Label in the first row of the results table indicates that it corresponds to the difference in the first two levels of A. Note that the estimated difference agrees with the value computed above. The difference is significant (p=0.0353) and a 95% large-sample confidence interval (0.01854, 0.5216) is also provided.
|
Binary Response — Using PROC NLMIXED
For details about using PROC NLMIXED, see the NLMIXED documentation (SAS Note 22930).
The differences in means can also be estimated by fitting the logistic model in PROC NLMIXED. With its ESTIMATE statement, you specify the nonlinear functions that estimate the differences. In PROC NLMIXED, you write the model on the event probability, p, and then specify p in the BINARY distribution option in the MODEL statement. The function LOGISTIC(x) = [1 + e-(x)]-1 makes it easy to specify the logistic model:
p = 1/(1+e-(Intercept + b1*A1 + b2*A2))
where A1 and A2 are two indicator variables (A1=1 if A=1, A1=0 otherwise; A2=1 if A=2, A2=0 otherwise). This is equivalent to reference parameterization (PARAM=REF) produced by the CLASS statement in several modeling procedures. The same model can be written in terms of the log odds (logit) as:
logit(p) = log(p/(1-p)) = Intercept + b1*A1 + b2*A2
Under this model, the estimated event probability for the A=1 population is 1/(1+e-(Intercept + b1)), for the A=2 population is 1/(1+e-(Intercept + b2)) and for the A=3 population is 1/(1+e-Intercept). The differences in probabilities between pairs of the three A populations are defined in the ESTIMATE statements. The LOGISTIC function is also used in the ESTIMATE statement to define the individual probabilities. Each ESTIMATE statement provides the estimated difference in probabilities along with its standard error and 95% confidence limits.
proc nlmixed data=test;
p=logistic(Intercept + b1*(a=1) + b2*(a=2));
model y ~ binary(p);
estimate "Prob Diff A1-A2" logistic(Intercept+b1) - logistic(Intercept+b2);
estimate "Prob Diff A1-A3" logistic(Intercept+b1) - logistic(Intercept);
estimate "Prob Diff A2-A3" logistic(Intercept+b2) - logistic(Intercept);
run;
The ESTIMATE statements produce the "Additional Estimates" table below. The estimated difference in probabilities between levels 1 and 2 of A is 0.2701 with a 95% confidence interval of (0.0155, 0.5247). Note these results, which use t-tests, closely match the large-sample results from the NLMeans macro.NOTE
|
|||||||||||||||||||||||||||||||||||||||||||||
Binary Response — Using the NLEstimate macro
See the description of the NLEstimate macro (SAS Note 58775) for information on making it available in your SAS® session.
The differences in probabilities can also be estimated with the NLEstimate macro. When fitting the model in PROC LOGISTIC, include the STORE statement to save the model. Create a data set with an observation for each function to be estimated. See the description of the NLEstimate macro for details.
These statements produce the same results as PROC NLMIXED above. Note that a large-sample (Wald) test and confidence interval can be obtained by simply removing the DF= option.NOTE
proc logistic data=test;
class a / param=glm;
model y(event="1") = a;
lsmeans a / ilink;
store logfit;
run;
data fd;
length label f $32767;
infile datalines delimiter=',';
input label f;
datalines;
Prob Diff A1-A2,logistic(B_p1+b_p2) - logistic(B_p1+b_p3)
Prob Diff A1-A3,logistic(B_p1+b_p2) - logistic(B_p1)
Prob Diff A2-A3,logistic(B_p1+b_p3) - logistic(B_p1)
;
%NLEstimate(instore=logfit, fdata=fd, df=100)
Binary Response — Using PROC GENMOD or PROC CATMOD
Another approach fits a linear probability model with PROC GENMOD (using maximum likelihood estimation) or PROC CATMOD (using weighted least squares estimation). Note that some data might not be well fit by a linear probability model. Various error conditions, such as invalid mean parameter (SAS Note 23001), can occur during model fitting. In PROC GENMOD, use the DIST=BINOMIAL and LINK=IDENTITY options to model the binomial probabilities directly, rather than the logits. For details about GENMOD, see the GENMOD documentation (SAS Note 22930).
proc genmod data=test descending;
class a;
model y = a / dist=binomial link=identity;
lsmeans a / diff cl;
run;
The "a Least Squares Means table" shows the estimated probabilities for A along with confidence intervals. The "Differences of a Least Squares Means" table provides the estimated differences in response probabilities. Confidence intervals for the differences are also provided. The estimated differences in probabilities closely match the NLMIXED results.NOTE
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The same linear probability model can be fit in PROC CATMOD. The RESPONSE statement specifies a linear combination of the ordered response probabilities, Pr(Y=0) and Pr(Y=1). So, the 0 1 specification causes CATMOD to model Pr(Y=1). Predictors are treated as categorical by default. The PARAM=REF option specifies reference parameterization for the resulting indicator variables. For details about CATMOD, see the CATMOD documentation (SAS Note 22930).
proc catmod data=test;
response 0 1;
model y = a / param=ref clparm;
contrast "Prob Diff A1-A2" a 1 -1 / estimate=parm;
contrast "Prob Diff A1-A3" a 1 0 / estimate=parm;
contrast "Prob Diff A2-A3" a 0 1 / estimate=parm;
run; quit;
The CONTRAST statements estimate the differences in probabilities for pairs of the levels of A as indicated in the labels. The estimated differences again closely match the results above.NOTE
|
||||||||||||||||||||||||||||||||||||||||||||||||||
Multinomial Response — Using PROC FREQ
The following data are described in the example titled "Nominal Response Data: Generalized Logits Model" in the PROC LOGISTIC documentation (SAS Note 22930). The three levels of Style are the levels of a multinomial response observed in each of three schools. The three schools are compared on their probabilities of choosing each style.
data School;
length Program $ 9;
input School Program $ Style $ Count @@;
datalines;
1 regular self 10 1 regular team 17 1 regular class 26
1 afternoon self 5 1 afternoon team 12 1 afternoon class 50
2 regular self 21 2 regular team 17 2 regular class 26
2 afternoon self 16 2 afternoon team 12 2 afternoon class 36
3 regular self 15 3 regular team 15 3 regular class 16
3 afternoon self 12 3 afternoon team 12 3 afternoon class 20
;
proc freq;
weight Count;
table School*Style / nopercent nocol;
run;
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
A nonmodeling approach can be taken using PROC FREQ to construct appropriate 2×2 subtables. The RISKDIFF option provides the estimate of the difference in probabilities along with a confidence interval. To construct each subtable, use formats to combine Styles (columns) and a WHERE statement to drop Schools (rows) as needed in each case.
proc format;
value $class "class"="1:class" "self","team" ="2:other";
value $self "self" ="1:self" "class","team"="2:other";
value $team "team" ="1:team" "class","self"="2:other";
run;
The following statements create and analyze a 2×2 table for the comparison of schools 1 and 2 on their probabilities of choosing each style. The WHERE statement in each case omits the data from school 3. Similar statements can be used to do comparisons of the other pairs of schools.
This table compares schools 1 and 2 on Pr(Style=class) after combining the self and team columns.
proc freq order=formatted;
where School ne 3;
format Style $class.;
weight Count;
table School*Style / riskdiff nopercent nocol;
run;
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
This table compares schools 1 and 2 on Pr(Style=self) after combining the class and team columns.
proc freq order=formatted;
where School ne 3;
format Style $self.;
weight Count;
table School*Style / riskdiff nopercent nocol;
run;
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
This table compares schools 1 and 2 on Pr(Style=team) after combining the class and self columns.
proc freq order=formatted;
where School ne 3;
format Style $team.;
weight Count;
table School*Style / riskdiff nopercent nocol;
run;
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
One criticism of this approach is that not all of the data are used in each analysis. The model-based analyses that follow avoid this problem. However, in this simple case in which School is treated as a nominal effect and there are no covariates, the results from a modeling approach matches the nonmodeling analysis.
Multinomial Response — Using the NLMeans macro
The following statements fit a logistic model, using generalized logits, to the nominal multinomial response. As with the binary example, the LSMEANS / ILINK E; statement provides estimated probabilities and displays the coefficients that define the LS-means. Estimated probabilities are provided for each School on the first two Style levels. The third Style level is not included in the results since that level is redundant given the first two by the constraint that the Style probabilities must sum to one.
proc logistic data=School;
freq Count;
class School / param=glm;
model Style(ref="team")=School / link=glogit;
lsmeans School / ilink e;
store out=MultMod;
ods output coef=Coeffs;
run;
This NLMeans macro call provides pairwise differences among the three Schools on all of the Style probabilities.
%NLMeans(instore=MultMod, coef=Coeffs, link=glogit, title=Differences of School Means)
As indicated by Label, the first three rows are the pairwise differences among the Schools on the first response probability (Style=class). The pairwise differences on the next two Style probabilities follow. The results for the School 1 vs. 2 difference agree with the PROC FREQ results above.
|
Multinomial Response — Using the NLEstimate macro
To estimate these differences with the NLEstimate macro, create a data set with an observation for each function to be estimated. See the description of the NLEstimate macro for details. These statements produce the same results as PROC NLMIXED below.
proc logistic data=School;
freq Count;
class School / param=glm;
model Style(ref="team")=School / noint link=glogit;
store out=MultMod;
run;
data fd;
length label f $32767;
infile datalines delimiter=',';
input label f;
datalines;
School 1 vs 2 on Pr(class),exp(b_p1)/(1+exp(b_p1)+exp(b_p2))-exp(b_p3)/(1+exp(b_p3)+exp(b_p4))
School 1 vs 2 on Pr(self),exp(b_p2)/(1+exp(b_p1)+exp(b_p2))-exp(b_p4)/(1+exp(b_p3)+exp(b_p4))
School 1 vs 2 on Pr(team),1/(1+exp(b_p1)+exp(b_p2))-1/(1+exp(b_p3)+exp(b_p4))
;
%NLEstimate(instore=MultMod, fdata=fd)
Multinomial Response — Using PROC CATMOD
As in the binary response case above, PROC CATMOD can fit a linear probability model using weighted least squares estimation. The linear combination of ordered response probabilities in the RESPONSE statement below selects the first two probabilities, Pr(Style=class) and Pr(Style=self), which are labeled as Function Number in the results. Again, only two of the three can be modeled since the constraint that they must sum to one limits the number of independent probabilities to two. The NOINT and PARAM=REF options cause the estimated parameters of the model to be the estimated probabilities for schools 1 and 2 on the first two probabilities.
proc catmod data=school;
weight Count;
response 1 0 0, 0 1 0;
model Style=School / clparm noint param=ref;
contrast 'School 1 vs 2 on Pr(class)' all_parms 1 0 -1 0 / estimate=parm;
contrast 'School 1 vs 2 on Pr(self)' all_parms 0 1 0 -1 / estimate=parm;
contrast 'School 1 vs 2 on Pr(team)' all_parms -1 -1 1 1 / estimate=parm;
run; quit;
The CONTRAST statements estimate the differences between Schools 1 and 2 on each Style. The ALL_PARMS keyword enables you to specify a list of multipliers that define a linear combination of model parameters. For example, in the first CONTRAST statement, the linear combination is (1)(0.6333)+(-1)(0.4844)=0.1490, which compares Schools 1 and 2 on Pr(Style=class). The second CONTRAST statement does similarly for the comparison on Pr(Style=self). The third CONTRAST statement estimates the difference on Pr(Style=team) using the fact that the sum of the Style probabilities within each school must sum to one. Similar sets of CONTRAST statements could be written to compare the other pairs of schools.
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Multinomial Response — Using PROC NLMIXED
PROC NLMIXED can fit a nominal, multinomial logistic model using maximum likelihood estimation. You can use its ESTIMATE statement to estimate differences in probabilities. However, since the multinomial distribution is not supported in the MODEL statement, it must be defined via the GENERAL distribution. The statements below define a generalized logit model:
log(OddsCT) = log(Pr(Style=class)/Pr(Style=team)) = βc1S1 + βc2S2 + βc3S3
log(OddsST) = log(Pr(Style=self)/Pr(Style=team)) = βs1S1 + βs2S2 + βs3S3
where S1=1 if School=1 and 0 otherwise, and similarly for S2 and S3. Under this model, the estimate of Pr(Style=team) in the School=1 population is 1/(1+OddsCT+OddsST), where OddsCT=exp(βc1) and OddsST=exp(βs1). The estimates for Pr(Style=class) and Pr(Style=self) simply multiply Pr(Style=team) by OddsCT or OddsST respectively.
The ESTIMATE statements produce estimates of the probability differences between Schools 1 and 2 on each Style. Similar ESTIMATE statements can be written to compare the other pairs of Schools.
proc nlmixed data=school;
/* Create numeric response variable */
if Style='class' then y=1;
else if Style='self' then y=2;
else y=3;
/* Specify the nominal multinomial logistic model */
OddsCT=exp( (School=1)*bc1 + (School=2)*bc2 + (School=3)*bc3 );
OddsST=exp( (School=1)*bs1 + (School=2)*bs2 + (School=3)*bs3 );
PrT=1/(1+OddsCT+OddsST);
PrS=PrT*OddsST;
PrC=PrT*OddsCT;
if y=1 then p=PrC;
else if y=2 then p=PrS;
else p=PrT;
/* Define multinomial log likelihood */
ll = Count*log(p);
model y ~ general(ll);
/* Compare schools 1 and 2 */
estimate 'School 1 vs 2 on Pr(class)' exp(bc1)/(1+exp(bc1)+exp(bs1))-exp(bc2)/(1+exp(bc2)+exp(bs2));
estimate 'School 1 vs 2 on Pr(self)' exp(bs1)/(1+exp(bc1)+exp(bs1))-exp(bs2)/(1+exp(bc2)+exp(bs2));
estimate 'School 1 vs 2 on Pr(team)' 1/(1+exp(bc1)+exp(bs1))-1/(1+exp(bc2)+exp(bs2));
run;
Estimates are the same as from the other methods.NOTE
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
_______
Note: The confidence limits produced by NLMIXED differ slightly from those produced by CATMOD, GENMOD, or the default (df= is not specified) from the NLMeans or NLEstimate macros because NLMIXED uses a quantile from the t distribution to multiply the standard error of the predicted value in forming the interval, while FREQ, GENMOD, and CATMOD use a quantile from the standard normal distribution. This difference diminishes as the sample size increases. To more closely match the results from the NLMeans or NLEstimate macros to those from NLMIXED, specify df= in the macro to use the same degrees of freedom as NLMIXED.
Operating System and Release Information
| Product Family | Product | System | SAS Release | |
| Reported | Fixed* | |||
| SAS System | SAS/STAT | z/OS | ||
| OpenVMS VAX | ||||
| Microsoft® Windows® for 64-Bit Itanium-based Systems | ||||
| Microsoft Windows Server 2003 Datacenter 64-bit Edition | ||||
| Microsoft Windows Server 2003 Enterprise 64-bit Edition | ||||
| Microsoft Windows XP 64-bit Edition | ||||
| Microsoft® Windows® for x64 | ||||
| OS/2 | ||||
| Microsoft Windows 95/98 | ||||
| Microsoft Windows 2000 Advanced Server | ||||
| Microsoft Windows 2000 Datacenter Server | ||||
| Microsoft Windows 2000 Server | ||||
| Microsoft Windows 2000 Professional | ||||
| Microsoft Windows NT Workstation | ||||
| Microsoft Windows Server 2003 Datacenter Edition | ||||
| Microsoft Windows Server 2003 Enterprise Edition | ||||
| Microsoft Windows Server 2003 Standard Edition | ||||
| Microsoft Windows Server 2008 | ||||
| Microsoft Windows XP Professional | ||||
| Windows Millennium Edition (Me) | ||||
| Windows Vista | ||||
| 64-bit Enabled AIX | ||||
| 64-bit Enabled HP-UX | ||||
| 64-bit Enabled Solaris | ||||
| ABI+ for Intel Architecture | ||||
| AIX | ||||
| HP-UX | ||||
| HP-UX IPF | ||||
| IRIX | ||||
| Linux | ||||
| Linux for x64 | ||||
| Linux on Itanium | ||||
| OpenVMS Alpha | ||||
| OpenVMS on HP Integrity | ||||
| Solaris | ||||
| Solaris for x64 | ||||
| Tru64 UNIX | ||||
| Type: | Usage Note |
| Priority: | |
| Topic: | SAS Reference ==> Procedures ==> CATMOD SAS Reference ==> Procedures ==> LOGISTIC SAS Reference ==> Procedures ==> NLMIXED Analytics ==> Categorical Data Analysis SAS Reference ==> Procedures ==> GENMOD SAS Reference ==> Macro |
| Date Modified: | 2022-10-14 14:19:03 |
| Date Created: | 2009-09-21 13:04:24 |