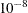-
ALPHA=

-
specifies the level  for the
for the 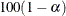 % confidence intervals for the survival functions. For example, ALPHA=0.05 requests 95% confidence limits. By default, ALPHA=0.05.
% confidence intervals for the survival functions. For example, ALPHA=0.05 requests 95% confidence limits. By default, ALPHA=0.05.
-
ALPHAQT=

-
specifies the level  for the
for the 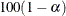 % confidence intervals for the quartiles of the survival time. For example, ALPHAQT=0.05 requests a 95% confidence interval.
By default, ALPHAQT=0.05.
% confidence intervals for the quartiles of the survival time. For example, ALPHAQT=0.05 requests a 95% confidence interval.
By default, ALPHAQT=0.05.
-
BOOTSTRAP <(options)>
BOOT <(options)>
-
performs simple bootstrap sampling and computes bootstrap standard errors of nonparametric survival estimates. You can specify
the following suboptions to control how the bootstrap samples are generated:
-
NBOOT
-
specifies the number of bootstrap samples to be generated. The default value is 1,000.
-
SEED
-
specifies the seed to generate bootstrap samples. The default seed is selected randomly.
-
CONFTYPE=ASINSQRT | LINEAR | LOG | LOGLOG
-
specifies the transformation to be applied to 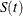 to obtain pointwise confidence intervals for the survival function in addition to the confidence intervals for the quartiles
of the survival times. You can specify the following keywords:
to obtain pointwise confidence intervals for the survival function in addition to the confidence intervals for the quartiles
of the survival times. You can specify the following keywords:
- ASINSQRT
-
specifies the arcsine–square root transformation,
- LOGLOG
-
specifies the log–log transformation,
This is also referred to as the log cumulative hazard transformation, because it applies the log transformation to the cumulative
hazard function. Collett (1994) and Lachin (2000) call it the complementary log-log transformation.
- LINEAR
-
specifies the identity transformation,
- LOG
-
specifies the log transformation,
- LOGIT
-
specifies the logit transformation,
By default, CONFTYPE=LOGLOG.
-
DATA=SAS-data-set
-
names the SAS data set that PROC ICLIFETEST reads. By default, the most recently created SAS data set is used.
-
IMPUTE <(options)>
IM <(options)>
-
specifies details for multiple imputations. You can specify the following two suboptions to control how the samples are generated:
-
NIMSE
-
specifies the number of imputation samples to be generated for computing standard errors of the survival estimates. The default
value is 1,000.
-
NIMTEST
-
specifies the number of imputation samples to be generated for estimating the covariance matrix of the generalized log-rank
statistic. The default value is 1,000.
-
SEED
-
specifies the seed to generate imputation data sets. The default seed is selected randomly.
-
ITHISTORY
-
prints the iteration history of the nonparametric estimation, including the log likelihood, the probability estimate that
is associated with each Turnbull interval, and whether the EM or ICM method is used for each iteration.
-
MAXITER=n
MAXIT=n
-
specifies the maximum number of iterations for estimating the survival function. The default value depends on the estimation
method as follows:
-
EMICM method: 200
-
ICM method: 200
-
Turnbull method: 500
Note that you specify the method with the METHOD= option.
-
MAXTIME=value
-
specifies the maximum value of the time variable that is allowed in plots so that outlying points do not determine the scale
of the time axis of the plots. This option affects only the way in which plots are displayed and has no effect on any calculations.
-
METHOD=TURNBULL | ICM | EMICM
-
specifies the method to be used for computing survival function estimates. You can specify the following:
-
TURNBULL
EM
-
requests that the Turnbull method be used to estimate the survival function.
-
EMICM
-
requests that the EMICM algorithm be used to estimate the survival function.
-
ICM
-
requests that the iterative convex minorant algorithm be used to estimate the survival function.
By default, METHOD=EMICM.
-
MISSING
-
allows missing values to be a stratum level or a valid group for the variables that are specified in the STRATA and TEST statements.
-
NOPRINT
-
suppresses the display of output. This option is useful when only an output data set is needed. It temporarily disables the
Output Delivery System (ODS); for more information about ODS, see Chapter 20: Using the Output Delivery System.
-
NOSUMMARY
-
suppresses the summary table of the number of censored and uncensored values.
-
OUTSURV=SAS-data-set
OUTS=SAS-data-set
-
creates an output SAS data set to contain the estimates of the survival function and corresponding confidence limits for all
strata. For more information about the contents of the OUTSURV= data set, see the section OUTSURV= Data Set.
-
PLOTS<(global-plot-options)>=plot-request <(options)>
PLOTS<(global-plot-options)>=(plot-request <(options)> <…plot-request <(options)>>)
-
controls the plots that are produced by using ODS Graphics. When you specify only one plot-request, you can omit the parentheses around it. Here are some examples:
plots=none
plots=(survival(cl) logsurv)
plots(only)=hazard
ODS Graphics must be enabled before plots can be requested. For example:
ods graphics on;
proc iclifetest plots=survival;
time (Left,Right);
run;
ods graphics off;
For more information about enabling and disabling ODS Graphics, see the section Enabling and Disabling ODS Graphics in Chapter 21: Statistical Graphics Using ODS.
If ODS Graphics is enabled but you do not specify the PLOTS= option, then by default PROC ICLIFETEST produces a plot of the
estimated survival functions.
You can specify the following global-plot-option:
-
ONLY
-
specifies that only the specified plots in the list be produced; otherwise, the default survival function plot is also displayed.
You can specify the following plot-requests and their options:
-
ALL
-
produces all appropriate plots. Specifying PLOTS=ALL is equivalent to specifying PLOTS=(SURVIVAL LOGSURV LOGLOGS HAZARD).
-
HAZARD <(hazard-options)>
H <hazard-options>
-
plots the Kernel-smoothed hazard functions.
You can specify the following hazard-options.
-
BANDWIDTH=value | numeric-list | RANGE(lower,upper)
BW=
-
specifies the bandwidth for kernel-smoothed estimation of the hazard function. You can specify one of the following:
-
value
-
sets the bandwidth to the given value.
-
numeric-list
-
selects the bandwidth from the specified numeric-list that minimizes the mean integrated squared error.
-
RANGE(lower,upper)
-
selects from the interval (lower, upper) the bandwidth that minimizes a cross validation pseudo-likelihood function. PROC ICLIFETEST uses the golden section search
algorithm to find the minimum. If the interval contains more than one local minimum, there is no guarantee that the local
minimum that the algorithm finds is also the global minimum.
For more information about the cross validation pseudo-likelihood function, see the section Kernel-Smoothed Estimation. By default, BANDWIDTH= RANGE(0.1b,0.25b), where 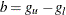 ;
; 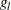 is the left boundary of the first Turnbull interval; and
is the left boundary of the first Turnbull interval; and 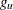 is the right boundary of the last Turnbull interval if it is finite or the left boundary of that interval multiplied by
is the right boundary of the last Turnbull interval if it is finite or the left boundary of that interval multiplied by 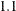 otherwise.
otherwise.
-
SAMPLING=LEAVEONE | RANDOM
-
specifies how to partition the data set to form cross validation groups. You can specify the following values:
-
LEAVEONE
-
partitions the data set into leave-one-out subsets.
-
RANDOM
-
randomly assigns observations to cross validation groups with equal probabilities.
By default, SAMPLING=RANDOM.
-
GRIDL=number
-
specifies the lower grid limit for the kernel-smoothed estimate. The default value is the time origin.
-
GRIDU=number
-
specifies the upper grid limit for the kernel-smoothed estimate. The default value is the maximum input boundary value.
-
KERNEL=kernel-option
-
specifies the kernel to be used. You can specify the following values:
-
BIWEIGHT
BW
-
uses the following kernel: 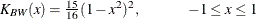
-
EPANECHNIKOV
E
-
uses the following kernel: 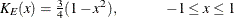
-
UNIFORM
U
-
uses the following kernel: 
By default, KERNEL=EPANECHNIKOV.
-
CVFOLD=number
-
specifies the number of cross validation groups. This option is applicable only when SAMPLING=RANDOM. The default number is either 5 or the sample size, whichever value is less.
-
CVGRID=number
-
specifies the number of grid points to use in determining the cross validation pseudo-likelihood. By default, CVGRID=51.
-
HGRID=number
-
specifies the number of grid points for discretizing the smoothed hazard function. By default, HGRID=101.
-
LOGLOGS
LLS
-
plots the log of the negative log of the estimated survival function versus the log of time.
-
LOGSURV
LS
-
plots the negative log of the estimated survival function versus time.
-
NONE
-
suppresses all plots.
-
SURVIVAL <(CL | FAILURE | NODASH | STRATA | TEST)>
S <(CL | FAILURE | STRATA | TEST)>
-
plots the estimated survival function. You can customize the display by specifying the following values:
-
CL
-
displays pointwise confidence limits for the survival function.
-
FAILURE
F
-
changes all the displays for survival function to those for the failure function. For example, if you specify both the FAILURE
and CL options, the plot displays the failure curves in addition to pointwise confidence limits for the failure function.
-
NODASH
-
suppresses the plotting of dashed lines as linear interpolations for the Turnbull intervals.
-
STRATA=INDIVIDUAL | OVERLAY | PANEL
-
specifies how to display the survival or failure curves for multiple strata. This option has no effect if there is only one
stratum. You can specify one of the following values:
-
INDIVIDUAL
UNPACK
-
requests that a separate plot be displayed for each stratum.
-
OVERLAY
-
requests that the survival or failure curves for the strata be overlaid on the same plot.
-
PANEL
-
requests that separate plots for the strata be organized into panels of two or four plots, depending on the number of strata.
By default, STRATA=OVERLAY.
-
TEST
-
displays the p-value for a K-sample test that is specified in the TEST statement.
-
SHOWTI
-
presents the survival estimates in terms of Turnbull intervals.
-
SINGULAR=value
-
sets the tolerance for testing the singularity of the covariance matrix of test statistics.
-
TOLLIKE=value
-
sets the log-likelihood convergence criterion for the estimation algorithm. The default value depends on the estimation method as follows:
-
EMICM method: 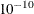
-
ICM method: 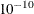
-
Turnbull method: