| The GENMOD Procedure |
Example 37.1 Logistic Regression
In an experiment comparing the effects of five different drugs, each drug is tested on a number of different subjects. The outcome of each experiment is the presence or absence of a positive response in a subject. The following artificial data represent the number of responses r in the n subjects for the five different drugs, labeled A through E. The response is measured for different levels of a continuous covariate x for each drug. The drug type and the continuous covariate x are explanatory variables in this experiment. The number of responses r is modeled as a binomial random variable for each combination of the explanatory variable values, with the binomial number of trials parameter equal to the number of subjects n and the binomial probability equal to the probability of a response.
The following DATA step creates the data set:
data drug; input drug$ x r n @@; datalines; A .1 1 10 A .23 2 12 A .67 1 9 B .2 3 13 B .3 4 15 B .45 5 16 B .78 5 13 C .04 0 10 C .15 0 11 C .56 1 12 C .7 2 12 D .34 5 10 D .6 5 9 D .7 8 10 E .2 12 20 E .34 15 20 E .56 13 15 E .8 17 20 ; run;
A logistic regression for these data is a generalized linear model with response equal to the binomial proportion r/n. The probability distribution is binomial, and the link function is logit. For these data, drug and x are explanatory variables. The probit and the complementary log-log link functions are also appropriate for binomial data.
PROC GENMOD performs a logistic regression on the data in the following SAS statements:
proc genmod data=drug;
class drug;
model r/n = x drug / dist = bin
link = logit
lrci;
run;
Since these data are binomial, you use the events/trials syntax to specify the response in the MODEL statement. Profile likelihood confidence intervals for the regression parameters are computed using the LRCI option.
General model and data information is produced in Output 37.1.1.
The five levels of the CLASS variable DRUG are displayed in Output 37.1.2.
In the "Criteria For Assessing Goodness Of Fit" table displayed in Output 37.1.3, the value of the deviance divided by its degrees of freedom is less than 1. A 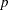 -value is not computed for the deviance; however, a deviance that is approximately equal to its degrees of freedom is a possible indication of a good model fit. Asymptotic distribution theory applies to binomial data as the number of binomial trials parameter n becomes large for each combination of explanatory variables. McCullagh and Nelder (1989) caution against the use of the deviance alone to assess model fit. The model fit for each observation should be assessed by examination of residuals. The OBSTATS option in the MODEL statement produces a table of residuals and other useful statistics for each observation.
-value is not computed for the deviance; however, a deviance that is approximately equal to its degrees of freedom is a possible indication of a good model fit. Asymptotic distribution theory applies to binomial data as the number of binomial trials parameter n becomes large for each combination of explanatory variables. McCullagh and Nelder (1989) caution against the use of the deviance alone to assess model fit. The model fit for each observation should be assessed by examination of residuals. The OBSTATS option in the MODEL statement produces a table of residuals and other useful statistics for each observation.
| Criteria For Assessing Goodness Of Fit | |||
|---|---|---|---|
| Criterion | DF | Value | Value/DF |
| Deviance | 12 | 5.2751 | 0.4396 |
| Scaled Deviance | 12 | 5.2751 | 0.4396 |
| Pearson Chi-Square | 12 | 4.5133 | 0.3761 |
| Scaled Pearson X2 | 12 | 4.5133 | 0.3761 |
| Log Likelihood | -114.7732 | ||
| Full Log Likelihood | -23.7343 | ||
| AIC (smaller is better) | 59.4686 | ||
| AICC (smaller is better) | 67.1050 | ||
| BIC (smaller is better) | 64.8109 | ||
In the "Analysis Of Parameter Estimates" table displayed in Output 37.1.4, chi-square values for the explanatory variables indicate that the parameter values other than the intercept term are all significant. The scale parameter is set to 1 for the binomial distribution. When you perform an overdispersion analysis, the value of the overdispersion parameter is indicated here. See the section Overdispersion for a discussion of overdispersion.
| Analysis Of Maximum Likelihood Parameter Estimates | ||||||||
|---|---|---|---|---|---|---|---|---|
| Parameter | DF | Estimate | Standard Error | Likelihood Ratio 95% Confidence Limits |
Wald Chi-Square | Pr > ChiSq | ||
| Intercept | 1 | 0.2792 | 0.4196 | -0.5336 | 1.1190 | 0.44 | 0.5057 | |
| x | 1 | 1.9794 | 0.7660 | 0.5038 | 3.5206 | 6.68 | 0.0098 | |
| drug | A | 1 | -2.8955 | 0.6092 | -4.2280 | -1.7909 | 22.59 | <.0001 |
| drug | B | 1 | -2.0162 | 0.4052 | -2.8375 | -1.2435 | 24.76 | <.0001 |
| drug | C | 1 | -3.7952 | 0.6655 | -5.3111 | -2.6261 | 32.53 | <.0001 |
| drug | D | 1 | -0.8548 | 0.4838 | -1.8072 | 0.1028 | 3.12 | 0.0773 |
| drug | E | 0 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | . | . |
| Scale | 0 | 1.0000 | 0.0000 | 1.0000 | 1.0000 | |||
| Note: | The scale parameter was held fixed. |
The preceding table contains the profile likelihood confidence intervals for the explanatory variable parameters requested with the LRCI option. Wald confidence intervals are displayed by default. Profile likelihood confidence intervals are considered to be more accurate than Wald intervals (see Aitkin et al. (1989)), especially with small sample sizes. You can specify the confidence coefficient with the ALPHA= option in the MODEL statement. The default value of 0.05, corresponding to 95% confidence limits, is used here. See the section Confidence Intervals for Parameters for a discussion of profile likelihood confidence intervals.
Copyright © SAS Institute, Inc. All Rights Reserved.