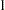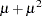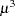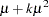| The GLIMMIX Procedure |
Implied Variance Functions
While link functions are not unique for each distribution (see Table 38.9 for the default link functions), the distribution does determine the variance function 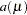 . This function expresses the variance of an observation as a function of the mean, apart from weights, frequencies, and additional scale parameters. The implied variance functions
. This function expresses the variance of an observation as a function of the mean, apart from weights, frequencies, and additional scale parameters. The implied variance functions 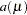 of the GLIMMIX procedure are shown in Table 38.15 for the supported distributions. For the binomial distribution,
of the GLIMMIX procedure are shown in Table 38.15 for the supported distributions. For the binomial distribution,  denotes the number of trials in the events/trials syntax. For the negative binomial distribution,
denotes the number of trials in the events/trials syntax. For the negative binomial distribution,  denotes the scale parameter. The multiplicative scale parameter
denotes the scale parameter. The multiplicative scale parameter 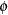 is not included for the other distributions. The last column of the table indicates whether
is not included for the other distributions. The last column of the table indicates whether 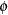 has a value equal to
has a value equal to  for the particular distribution.
for the particular distribution.
Variance function |
|||
|---|---|---|---|
DIST= |
Distribution |
|
|
BETA |
beta |
|
No |
BINARY |
binary |
|
Yes |
BINOMIAL |
binomial |
|
Yes |
EXPONENTIAL |
exponential |
|
Yes |
GAMMA |
gamma |
|
No |
GAUSSIAN |
normal |
|
No |
GEOMETRIC |
geometric |
|
Yes |
INVGAUSS |
inverse gaussian |
|
No |
LOGNORMAL |
lognormal |
|
No |
NEGBINOMIAL |
negative binomial |
|
Yes |
POISSON |
Poisson |
|
Yes |
TCENTRAL |
|
|
No |
To change the variance function, you can use SAS programming statements and the predefined automatic variables, as outlined in the following section. Your definition of a variance function will override the DIST= option and its implied variance function. This has the following implication for parameter estimation with the GLIMMIX procedure. When a user-defined link is available, the distribution of the data is determined from the DIST= option, or the respective default for the type of response. In a GLM, for example, this enables maximum likelihood estimation. If a user-defined variance function is provided, the DIST= option is not honored and the distribution of the data is assumed unknown. In a GLM framework, only quasi-likelihood estimation is then available to estimate the model parameters.
Copyright © 2009 by SAS Institute Inc., Cary, NC, USA. All rights reserved.
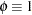
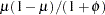
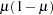
 |
|