| Language Reference |
MCD Call
finds the minimum covariance determinant estimator
- CALL MCD( sc, coef, dist, opt,
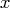 );
);
The MCD call is the robust (resistant) estimation of multivariate location and scatter, defined by minimizing the determinant of the covariance matrix computed from
The MCD subroutine computes the minimum covariance determinant estimator. These robust locations and covariance matrices can be used to detect multivariate outliers and leverage points. For this purpose, the MCD subroutine provides a table of robust distances.
In the following discussion,
- opt
- refers to an options vector with the following components
(missing values are treated as default values):
- opt[1]
- specifies the amount of printed output.
Higher option values request additional
output and include the output of lower values.
- opt[1]=0
- prints no output except error messages.
- opt[1]=1
- prints most of the output.
- opt[1]=2
- additionally prints case numbers of the
observations in the best subset and some
basic history of the optimization process.
- opt[1]=3
- additionally prints how many subsets result in singular linear systems.
The default is opt[1]=0.
- opt[2]
- specifies whether the classical, initial, and
final robust covariance matrices are printed.
The default is opt[2]=0.
Note that the final robust covariance
matrix is always returned in coef.
- opt[3]
- specifies whether the classical, initial, and final
robust correlation matrices are printed or returned:
- opt[3]=0
- does not return or print.
- opt[3]=1
- prints the robust correlation matrix.
- opt[3]=2
- returns the final robust correlation matrix in coef.
- opt[3]=3
- prints and returns the final robust correlation matrix.
- opt[4]
- specifies the quantile
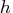 used in the objective function.
The default is opt[4]=
used in the objective function.
The default is opt[4]= ![h = [\frac{n+n+1}2]](images/langref_langrefeq760.gif) .
If the value of
.
If the value of 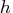 is specified outside the range
is specified outside the range 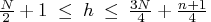 ,
it is reset to the closest boundary of this region.
,
it is reset to the closest boundary of this region.
- opt[5]
- specifies the number
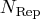 of subset generations.
This option is the same as described
for the LTS subroutines.
Due to computer time restrictions, not all subset combinations
can be inspected for larger values of
of subset generations.
This option is the same as described
for the LTS subroutines.
Due to computer time restrictions, not all subset combinations
can be inspected for larger values of 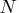 and
and 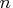 .
.
When opt[5] is zero or missing:
If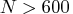 , construct up to five disjoint random subsets
with sizes as equal as possible, but not to exceed 300.
Inside each subset, choose
, construct up to five disjoint random subsets
with sizes as equal as possible, but not to exceed 300.
Inside each subset, choose 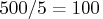 subset
combinations of
subset
combinations of 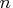 observations.
observations.
If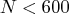 , the number of subsets is taken from the
following table.
, the number of subsets is taken from the
following table.
| n | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
| 500 | 50 | 22 | 17 | 15 | 14 | 0 | 0 | 0 | 0 |
| n | 11 | 12 | 13 | 14 | 15 |
| 0 | 0 | 0 | 0 | 0 |
- If the number of cases (observations)
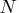 is smaller
than
is smaller
than 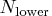 , then all possible subsets are used;
otherwise, 500 subsets are chosen randomly.
This means that an exhaustive search
is performed for opt[5]=-1.
If
, then all possible subsets are used;
otherwise, 500 subsets are chosen randomly.
This means that an exhaustive search
is performed for opt[5]=-1.
If 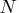 is larger than
is larger than 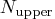 , a note is printed
in the log file indicating how many subsets exist.
, a note is printed
in the log file indicating how many subsets exist.
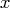
- refers to an
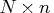 matrix
matrix 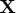 of regressors.
of regressors.
Missing values are not permitted in
The MCD subroutine returns the following values:
- sc
- is a column vector containing the following scalar information:
- sc[1]
- the quantile
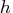 used in the objective function
used in the objective function
- sc[2]
- number of subsets generated
- sc[3]
- number of subsets with singular linear systems
- sc[4]
- number of nonzero weights
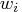
- sc[5]
- lowest value of the objective function
 attained (smallest determinant)
attained (smallest determinant)
- sc[6]
- Mahalanobis-like distance used in the computation of
the lowest value of the objective function

- sc[7]
- the cutoff value used for the outlier decision
- coef
- is a matrix with
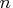 columns containing
the following results in its rows:
columns containing
the following results in its rows:
- coef[1]
- location of ellipsoid center
- coef[2]
- eigenvalues of final robust scatter matrix
- coef[3:2+n]
- the final robust scatter matrix for
opt[2]=1 or opt[2]=3
- coef[2+n+1:2+2n]
- the final robust correlation matrix for
opt[3]=1 or opt[3]=3
- dist
- is a matrix with
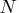 columns containing
the following results in its rows:
columns containing
the following results in its rows:
- dist[1]
- Mahalanobis distances
- dist[2]
- robust distances based on the final estimates
- dist[3]
- weights (=1 for small, =0 for large robust distances)
Example
Consider Brownlee's (1965) stackloss data used in the example for the MVE subroutine.For ![]() and
and ![]() (three explanatory variables
including intercept), you obtain a total of 5,985
different subsets of 4 observations out of 21.
If you decide not to specify optn[5],
the MCD algorithm chooses 500 random sample subsets,
as in the following code:
(three explanatory variables
including intercept), you obtain a total of 5,985
different subsets of 4 observations out of 21.
If you decide not to specify optn[5],
the MCD algorithm chooses 500 random sample subsets,
as in the following code:
/* X1 X2 X3 Y Stackloss data */
aa = { 1 80 27 89 42,
1 80 27 88 37,
1 75 25 90 37,
1 62 24 87 28,
1 62 22 87 18,
1 62 23 87 18,
1 62 24 93 19,
1 62 24 93 20,
1 58 23 87 15,
1 58 18 80 14,
1 58 18 89 14,
1 58 17 88 13,
1 58 18 82 11,
1 58 19 93 12,
1 50 18 89 8,
1 50 18 86 7,
1 50 19 72 8,
1 50 19 79 8,
1 50 20 80 9,
1 56 20 82 15,
1 70 20 91 15 };
a = aa[,2:4]; optn = j(8,1,.); optn[1]= 2; /* ipri */ optn[2]= 1; /* pcov: print COV */ optn[3]= 1; /* pcor: print CORR */ CALL MCD(sc,xmcd,dist,optn,a);
The first part of the output of this program
is a summary of the MCD algorithm
and the final ![]() points selected, as follows:
points selected, as follows:
Fast MCD by Rousseeuw and Van Driessen
Number of Variables 3
Number of Observations 21
Default Value for h 12
Specified Value for h 12
Breakdown Value 42.86
- Highest Possible Breakdown Value -
The best half of the entire data set obtained after full
iteration consists of the cases:
4 5 6 7 8 9 10 11 12 13 14 20
The second part of the output is the MCD estimators of the location, scatter matrix, and correlation matrix, as follows:
MCD Location Estimate
VAR1 VAR2 VAR3
59.5 20.833333333 87.333333333
Average of 12 Selected Points
MCD Scatter Matrix Estimate
VAR1 VAR2 VAR3
VAR1 5.1818181818 4.8181818182 4.7272727273
VAR2 4.8181818182 7.6060606061 5.0606060606
VAR3 4.7272727273 5.0606060606 19.151515152
Determinant = 238.07387929
Covariance Matrix of 12 Selected Points
MCD Correlation Matrix
VAR1 VAR2 VAR3
VAR1 1 0.7674714142 0.4745347313
VAR2 0.7674714142 1 0.4192963398
VAR3 0.4745347313 0.4192963398 1
The MCD scatter matrix is multiplied by a factor to make it
consistent when all the data come from a single Gaussian
distribution.
Consistent Scatter Matrix
VAR1 VAR2 VAR3
VAR1 8.6578437815 8.0502757968 7.8983838007
VAR2 8.0502757968 12.708297013 8.4553211199
VAR3 7.8983838007 8.4553211199 31.998580526
Determinant = 397.77668436
The final output presents a table containing the classical
Mahalanobis distances, the robust distances, and the weights
identifying the outlying observations (that is, leverage points
when explaining ![]() with these three regressor variables):
with these three regressor variables):
Classical Distances and Robust (Rousseeuw) Distances
Unsquared Mahalanobis Distance and
Unsquared Rousseeuw Distance of Each Observation
Mahalanobis Robust
N Distances Distances Weight
1 2.253603 12.173282 0
2 2.324745 12.255677 0
3 1.593712 9.263990 0
4 1.271898 1.401368 1.000000
5 0.303357 1.420020 1.000000
6 0.772895 1.291188 1.000000
7 1.852661 1.460370 1.000000
8 1.852661 1.460370 1.000000
9 1.360622 2.120590 1.000000
10 1.745997 1.809708 1.000000
11 1.465702 1.362278 1.000000
12 1.841504 1.667437 1.000000
13 1.482649 1.416724 1.000000
14 1.778785 1.988240 1.000000
15 1.690241 5.874858 0
16 1.291934 5.606157 0
17 2.700016 6.133319 0
18 1.503155 5.760432 0
19 1.593221 6.156248 0
20 0.807054 2.172300 1.000000
21 2.176761 7.622769 0
Robust distances are based on reweighted estimates.
The cutoff value is the square root of the 0.975 quantile of
the chi square distribution with 3 degrees of freedom.
Points whose robust distance exceeds 3.0575159206 have received
a zero weight in the last column above.
There were 9 such points in the data.
These may include boundary cases.
Only points whose robust distance is substantially larger
than the cutoff should be considered outliers.
Copyright © 2009 by SAS Institute Inc., Cary, NC, USA. All rights reserved.