The GLMPOWER Procedure
The univariate linear model has the form
where ![]() is the
is the ![]() vector of responses,
vector of responses, ![]() is the
is the ![]() design matrix,
design matrix, ![]() is the
is the ![]() vector of model parameters corresponding to the columns of
vector of model parameters corresponding to the columns of ![]() , and
, and ![]() is an
is an ![]() vector of errors with
vector of errors with
In PROC GLMPOWER, the model parameters ![]() are not specified directly, but rather indirectly as
are not specified directly, but rather indirectly as ![]() , which represents either conjectured response means or typical response values for each design profile. The
, which represents either conjectured response means or typical response values for each design profile. The ![]() values are manifested as the dependent variable in the MODEL statement. The vector
values are manifested as the dependent variable in the MODEL statement. The vector ![]() is obtained from
is obtained from ![]() according to the least squares equation,
according to the least squares equation,
Note that, in general, there is not a 1-to-1 mapping between ![]() and
and ![]() . Many different scenarios for
. Many different scenarios for ![]() might lead to the same
might lead to the same ![]() . If you specify
. If you specify ![]() with the intention of representing cell means, keep in mind that PROC GLMPOWER allows scenarios that are not valid cell means according to the model specified in the MODEL statement. For example, if
with the intention of representing cell means, keep in mind that PROC GLMPOWER allows scenarios that are not valid cell means according to the model specified in the MODEL statement. For example, if ![]() exhibits an interaction effect but the corresponding interaction term is left out of the model, then the cell means (
exhibits an interaction effect but the corresponding interaction term is left out of the model, then the cell means (![]() ) derived from
) derived from ![]() differ from
differ from ![]() . In particular, the cell means thus derived are the projection of
. In particular, the cell means thus derived are the projection of ![]() onto the model space.
onto the model space.
It is convenient in power analysis to parameterize the design matrix ![]() in three parts,
in three parts, ![]() , defined as follows:
, defined as follows:
-
The
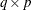 essence design matrix
essence design matrix 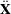 is the collection of unique rows of
is the collection of unique rows of 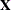 . Its rows are sometimes referred to as “design profiles.” Here,
. Its rows are sometimes referred to as “design profiles.” Here, 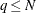 is defined simply as the number of unique rows of
is defined simply as the number of unique rows of 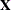 .
.
-
The
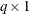 weight vector
weight vector 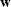 reveals the relative proportions of design profiles. Row i of
reveals the relative proportions of design profiles. Row i of 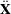 is to be included in the design
is to be included in the design 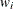 times for every
times for every 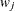 times row j is included. The weights are assumed to be standardized (that is, sum up to 1).
times row j is included. The weights are assumed to be standardized (that is, sum up to 1).
-
The total sample size is N. This is the number of rows in
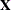 . If you gather
. If you gather 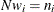 copies of the
copies of the 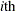 row of
row of 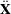 , for
, for 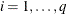 , then you end up with
, then you end up with 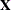 .
.
It is useful to express the crossproduct matrix ![]() in terms of these three parts,
in terms of these three parts,
since this factors out the portion (N) depending on sample size and the portion (![]() ) depending only on the design structure.
) depending only on the design structure.
A general linear hypothesis for the univariate model has the form
|
|
|
|
|
|
where ![]() is an
is an ![]() contrast matrix (assumed to be full rank) and
contrast matrix (assumed to be full rank) and ![]() is the null value (usually just a vector of zeros). Note that effect tests are just contrasts that use special forms of
is the null value (usually just a vector of zeros). Note that effect tests are just contrasts that use special forms of ![]() . Thus, this scheme covers both effect tests and custom contrasts.
. Thus, this scheme covers both effect tests and custom contrasts.
The test statistic is
where
|
|
|
|
|
|
|
|
|
where ![]() . Note that
. Note that ![]() if
if ![]() has full rank.
has full rank.
Under ![]() ,
, ![]() . Under
. Under ![]() , F is distributed as
, F is distributed as ![]() with noncentrality
with noncentrality
Muller and Peterson (1984) give the exact power of the test as
Sample size is computed by inverting the power equation.
See Muller and Benignus (1992) and O’Brien and Shieh (1992) for additional discussion.
