| The ALLELE Procedure |
Frequency Estimates
A marker locus M can have a series of alleles  ,
,  . A sample of
. A sample of  individuals can therefore have several different genotypes at the locus, with
individuals can therefore have several different genotypes at the locus, with  copies of type
copies of type  . The number
. The number  of copies of allele
of copies of allele  can be found directly by summation:
can be found directly by summation: 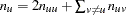 . The sample frequencies are written as
. The sample frequencies are written as  and
and  . The
. The  ’s are unbiased maximum likelihood estimates (MLEs) of the population proportions
’s are unbiased maximum likelihood estimates (MLEs) of the population proportions  .
.
The variance of the sample allele frequency 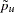 is calculated as
is calculated as
 |
and can be estimated by replacing  and
and  with their sample values
with their sample values 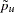 and
and  . The variance of the sample genotype frequency
. The variance of the sample genotype frequency  is not generally calculated; instead, an MLE of the HWD coefficient
is not generally calculated; instead, an MLE of the HWD coefficient 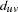 for alleles
for alleles  and
and  is calculated as
is calculated as
 |
and the MLE’s variance is estimated using one of the following formulas, depending on whether the two alleles are the same or different:
 |
 |
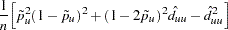 |
|||
 |
 |
 |
|||
 |
 |
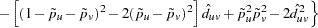 |
The standard error, the square root of the variance, is reported for the sample allele frequencies and the disequilibrium coefficient estimates. When the BOOTSTRAP= option of the PROC ALLELE statement is specified, bootstrap confidence intervals are formed by resampling individuals from the data set and are reported for these estimates, with the  % confidence level given by the ALPHA=
% confidence level given by the ALPHA= option (or
option (or  by default).
by default).
Copyright © 2008 by SAS Institute Inc., Cary, NC, USA. All rights reserved.