| The GENMOD Procedure |
| LSMEANS Statement |
- LSMEANS effects </ options> ;
The LSMEANS statement computes least squares means (LS-means) corresponding to the specified effects for the linear predictor part of the model. The  matrix constructed to compute them is precisely the same as the one formed in PROC GLM.
matrix constructed to compute them is precisely the same as the one formed in PROC GLM.
The LSMEANS statement is not available for multinomial distribution models for ordinal response data.
Each LS-mean is computed as 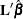 , where
, where  is the coefficient matrix associated with the least squares mean and
is the coefficient matrix associated with the least squares mean and 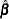 is the estimate of the parameter vector. The approximate standard errors for the LS-mean is computed as the square root of
is the estimate of the parameter vector. The approximate standard errors for the LS-mean is computed as the square root of 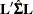 , where
, where 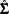 is the estimated covariance matrix of the parameter estimates. If you specify a GEE model in the REPEATED statement,
is the estimated covariance matrix of the parameter estimates. If you specify a GEE model in the REPEATED statement,  is the empirical covariance matrix estimate.
is the empirical covariance matrix estimate.
LS-means can be computed for any effect in the MODEL statement that involves CLASS variables. You can specify multiple effects in one LSMEANS statement or multiple LSMEANS statements, and all LSMEANS statements must appear after the MODEL statement.
As in the ESTIMATE statement, the  matrix is tested for estimability, and if this test fails, PROC GENMOD displays "Non-est" for the LS-means entries.
matrix is tested for estimability, and if this test fails, PROC GENMOD displays "Non-est" for the LS-means entries.
Assuming the LS-mean is estimable, PROC GENMOD constructs a Wald chi-square test to test the null hypothesis that the associated population quantity equals zero.
You can specify the following options in the LSMEANS statement after a slash (/).
- ALPHA=number
requests that a confidence interval be constructed for each of the LS-means with confidence level
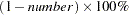 . The value of number must be between 0 and 1; the default value is 0.05, corresponding to a 95% confidence interval.
. The value of number must be between 0 and 1; the default value is 0.05, corresponding to a 95% confidence interval. - CL
requests that confidence limits be constructed for each of the LS-means. The confidence level is 0.95 by default; this can be changed with the ALPHA= option.
- CORR
displays the estimated correlation matrix of the LS-means as part of the "Least Squares Means" table.
- COV
displays the estimated covariance matrix of the LS-means as part of the "Least Squares Means" table.
- DIFF
requests that differences of the LS-means be displayed. All possible differences of LS-means, standard errors, and a Wald chi-square test are computed. Confidence limits are computed if the CL option is also specified.
- E
requests that the
 matrix coefficients for all LSMEANS effects be displayed.
matrix coefficients for all LSMEANS effects be displayed. - SINGULAR=number
- EPSILON=number
tunes the estimability checking as described for the CONTRAST statement.
Copyright © 2009 by SAS Institute Inc., Cary, NC, USA. All rights reserved.