| The ALLELE Procedure |
| Missing Values |
An individual’s genotype for a marker is considered missing if at least one of the alleles at the marker is missing. Any missing genotypes are excluded from all calculations, including the linkage disequilibrium statistics for all pairs that include the marker. However, the individual’s nonmissing genotypes at other markers can be used as part of the calculations.
If the BOOTSTRAP= option is specified, any individuals with missing genotypes for all markers are excluded from resampling. All other individuals are included, which could result in different numbers of individuals with nonmissing genotypes for the same marker across different samples.
When the POP statement is used, individuals with a missing value for the variable specified are excluded from the calculation of the population 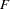 statistics, but they are included in all other analyses.
statistics, but they are included in all other analyses.
Copyright © 2008 by SAS Institute Inc., Cary, NC, USA. All rights reserved.